5.1.6 Scikit-learn and Matplotlib Hands-on Workshop
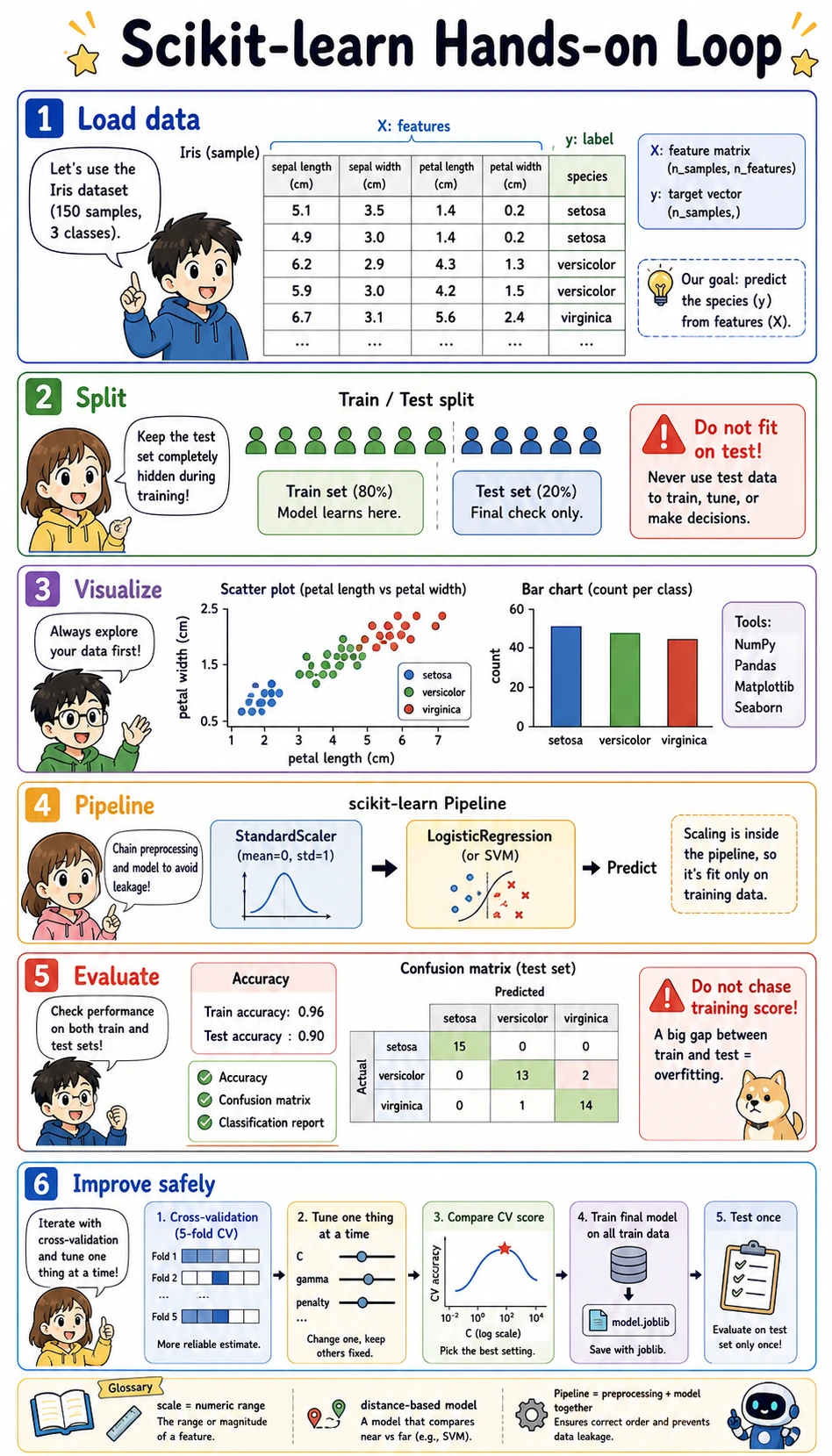

This is a follow-along workshop. The goal is not to add more theory, but to help you run a complete classic machine learning workflow by yourself: load data, visualize first, split data, train a model, evaluate it, improve it safely, and save it.
Learning objectives
- Understand what
X,y,X_train,X_test,y_train, andy_testmean in real code - Use Matplotlib to read data and model results before trusting a score
- Build a sklearn
Pipelinethat combines preprocessing and a model - Compare training and test scores without being fooled by overfitting
- Use cross-validation to tune one thing at a time
- Save and reload a trained Pipeline with
joblib
Prepare one runnable cell
Create a new notebook or Python file, then run this setup first.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import load_wine
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import ConfusionMatrixDisplay, classification_report
np.set_printoptions(precision=3, suppress=True)
If import sklearn fails, install the packages in the same Python environment:
python -m pip install --upgrade scikit-learn matplotlib joblib
pip installs packages. python -m pip means “use the pip that belongs to this exact Python interpreter,” which avoids the common mistake of installing into one environment and running code in another.
Load data: separate features and labels
In sklearn examples, you will see X and y all the time:
Xis the feature matrix. Each row is one sample, and each column is one input feature.yis the target vector. Each value is the answer label we want the model to learn.X.shapetells you(number_of_samples, number_of_features).y.shapetells you how many labels you have.
wine = load_wine()
X = wine.data
y = wine.target
print("X shape:", X.shape)
print("y shape:", y.shape)
print("Feature names:", wine.feature_names[:5], "...")
print("Class names:", wine.target_names.tolist())
print("First sample features:", np.round(X[0], 2))
print("First sample label:", y[0], "=>", wine.target_names[y[0]])
Expected output:
X shape: (178, 13)
y shape: (178,)
Feature names: ['alcohol', 'malic_acid', 'ash', 'alcalinity_of_ash', 'magnesium'] ...
Class names: ['class_0', 'class_1', 'class_2']
First sample features: [ 14.23 1.71 2.43 15.6 127. 2.8 3.06 0.28 2.29 5.64 1.04 3.92 1065. ]
First sample label: 0 => class_0
Before training any model, always answer three questions: “What is one row? What is one column? What is the label?” If those are unclear, the score will not mean much yet.
Matplotlib basics: read the chart before the model
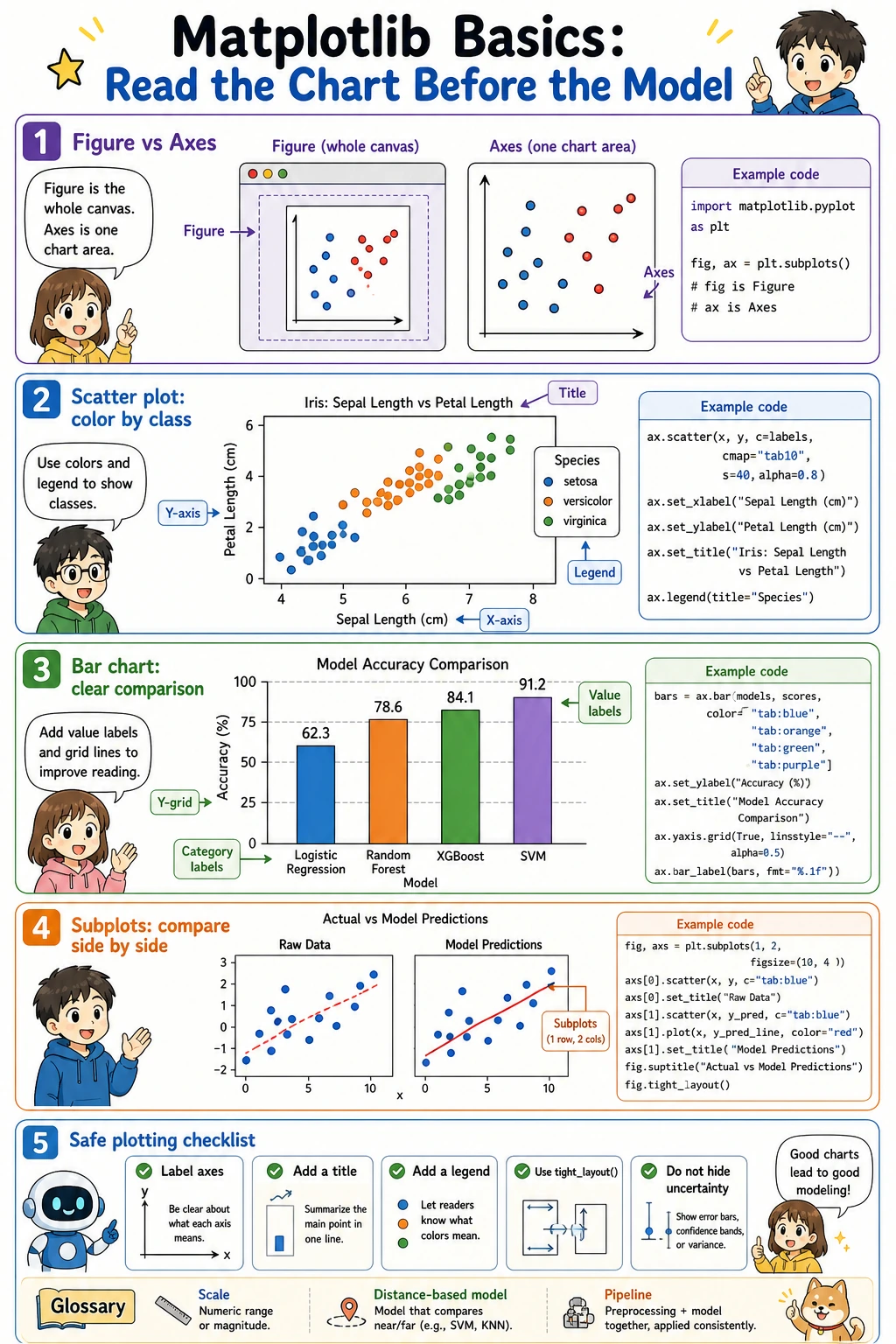
Matplotlib has two words that confuse beginners:
Figure: the whole canvas.Axes: one chart area inside the canvas.
Most beginner code can follow this pattern:
fig, ax = plt.subplots(figsize=(6, 4))
ax.scatter(x_values, y_values)
ax.set_xlabel("x-axis label")
ax.set_ylabel("y-axis label")
ax.set_title("Chart title")
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
Now draw two Wine features:
feature_x = 0 # alcohol
feature_y = 6 # flavanoids
fig, ax = plt.subplots(figsize=(7, 5))
scatter = ax.scatter(
X[:, feature_x],
X[:, feature_y],
c=y,
cmap="viridis",
s=45,
alpha=0.85,
)
ax.set_xlabel(wine.feature_names[feature_x])
ax.set_ylabel(wine.feature_names[feature_y])
ax.set_title("Wine data: two-feature view")
ax.grid(True, alpha=0.3)
ax.legend(
handles=scatter.legend_elements()[0],
labels=wine.target_names.tolist(),
title="Class",
)
plt.tight_layout()
plt.show()
What to observe:
- Are the classes already somewhat separated?
- Are there overlapping regions?
- Does one feature have a much larger numeric range than another feature?
This is why visualization matters: it gives you a first feeling for whether the model is solving an easy or difficult problem.
Split data: keep the test set hidden
train_test_split creates a training set and a test set.
- Training set: the model is allowed to learn from it.
- Test set: the model should only see it at the final evaluation step.
stratify=y: keep class proportions similar in train and test.random_state: make the split reproducible.
X_train, X_test, y_train, y_test = train_test_split(
X,
y,
test_size=0.2,
random_state=42,
stratify=y,
)
print("X_train:", X_train.shape, "y_train:", y_train.shape)
print("X_test: ", X_test.shape, "y_test: ", y_test.shape)
Expected output:
X_train: (142, 13) y_train: (142,)
X_test: (36, 13) y_test: (36,)
Do not run fit on the test set. The test set is your final exam. If preprocessing or tuning learns from the test set, the reported score becomes too optimistic.
Build a Pipeline: preprocessing plus model
Many models, such as logistic regression, SVM, and KNN, are sensitive to feature scale. The Wine dataset has columns with very different units, so we put StandardScaler before the model.
model = make_pipeline(
StandardScaler(),
LogisticRegression(max_iter=1000, random_state=42),
)
model.fit(X_train, y_train)
train_score = model.score(X_train, y_train)
test_score = model.score(X_test, y_test)
print(f"Train accuracy: {train_score:.1%}")
print(f"Test accuracy: {test_score:.1%}")
Expected output:
Train accuracy: 100.0%
Test accuracy: 100.0%
Pipeline matters because it keeps the correct order:
- On training data:
StandardScaler.fit_transformthen modelfit - On test data:
StandardScaler.transformthen modelpredict
That tiny difference prevents data leakage.
Predict and inspect concrete examples
A score is useful, but beginners should also look at a few actual predictions.
y_pred = model.predict(X_test)
proba = model.predict_proba(X_test[:5])
for i in range(5):
predicted_name = wine.target_names[y_pred[i]]
true_name = wine.target_names[y_test[i]]
confidence = proba[i].max()
print(f"Sample {i}: predicted={predicted_name}, true={true_name}, confidence={confidence:.1%}")
Example output:
Sample 0: predicted=class_0, true=class_0, confidence=99.9%
Sample 1: predicted=class_1, true=class_1, confidence=99.9%
Sample 2: predicted=class_0, true=class_0, confidence=99.5%
Sample 3: predicted=class_1, true=class_1, confidence=99.7%
Sample 4: predicted=class_2, true=class_2, confidence=99.9%
predict returns the final class. predict_proba returns the probability distribution over classes. Probability is useful when a business process needs thresholds, manual review, or risk ranking.
Evaluate with a confusion matrix and report
Accuracy alone hides which classes are confused with each other. A confusion matrix shows actual labels on one axis and predicted labels on the other axis.
fig, ax = plt.subplots(figsize=(5, 5))
ConfusionMatrixDisplay.from_estimator(
model,
X_test,
y_test,
display_labels=wine.target_names,
cmap="Blues",
ax=ax,
colorbar=False,
)
ax.set_title("Confusion matrix on test set")
plt.tight_layout()
plt.show()
print(classification_report(y_test, y_pred, target_names=wine.target_names))
What to read:
- Diagonal cells are correct predictions.
- Off-diagonal cells are mistakes.
- Precision asks: “Among predicted class A, how many were really A?”
- Recall asks: “Among real class A, how many did we catch?”
- F1 combines precision and recall.
Compare several models with the same workflow
Because sklearn has a unified API, model comparison is very practical.
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
models = {
"Logistic Regression": make_pipeline(
StandardScaler(),
LogisticRegression(max_iter=1000, random_state=42),
),
"Decision Tree": DecisionTreeClassifier(max_depth=4, random_state=42),
"KNN": make_pipeline(StandardScaler(), KNeighborsClassifier(n_neighbors=5)),
"SVM": make_pipeline(StandardScaler(), SVC(kernel="rbf", C=1.0, gamma="scale")),
}
results = {}
for name, clf in models.items():
clf.fit(X_train, y_train)
results[name] = {
"train": clf.score(X_train, y_train),
"test": clf.score(X_test, y_test),
}
print(f"{name:20s} train={results[name]['train']:.1%} test={results[name]['test']:.1%}")
Example output:
Logistic Regression train=100.0% test=100.0%
Decision Tree train=99.3% test=94.4%
KNN train=97.9% test=97.2%
SVM train=100.0% test=100.0%
Now draw the comparison:
fig, ax = plt.subplots(figsize=(9, 5))
names = list(results.keys())
x = np.arange(len(names))
width = 0.35
train_scores = [results[name]["train"] for name in names]
test_scores = [results[name]["test"] for name in names]
bars_train = ax.bar(x - width / 2, train_scores, width, label="Train", color="steelblue")
bars_test = ax.bar(x + width / 2, test_scores, width, label="Test", color="coral")
ax.set_xticks(x)
ax.set_xticklabels(names, rotation=15, ha="right")
ax.set_ylabel("Accuracy")
ax.set_title("Model comparison on Wine dataset")
ax.set_ylim(0.8, 1.05)
ax.legend()
ax.grid(axis="y", alpha=0.3)
ax.bar_label(bars_train, fmt="%.2f", padding=3)
ax.bar_label(bars_test, fmt="%.2f", padding=3)
plt.tight_layout()
plt.show()
If train score is much higher than test score, suspect overfitting. If both scores are low, suspect underfitting, weak features, or an unsuitable model.
Tune safely with cross-validation
Do not tune hyperparameters directly on the test set. Use cross-validation on the training set.
candidates = [0.01, 0.1, 1.0, 10.0, 100.0]
for C in candidates:
clf = make_pipeline(
StandardScaler(),
LogisticRegression(C=C, max_iter=1000, random_state=42),
)
scores = cross_val_score(clf, X_train, y_train, cv=5, scoring="accuracy")
print(f"C={C:<6} CV accuracy={scores.mean():.1%} ± {scores.std():.1%}")
Example output:
C=0.01 CV accuracy=95.8% ± 3.1%
C=0.1 CV accuracy=98.6% ± 1.8%
C=1.0 CV accuracy=98.6% ± 1.8%
C=10.0 CV accuracy=97.9% ± 2.6%
C=100.0 CV accuracy=97.9% ± 2.6%
The habit is more important than this exact result:
- Split off a test set and do not touch it.
- Tune with cross-validation on the training set.
- Choose the best setting.
- Train one final model on all training data.
- Evaluate on the test set once.
Save and reload the final Pipeline
import joblib
final_model = make_pipeline(
StandardScaler(),
LogisticRegression(C=1.0, max_iter=1000, random_state=42),
)
final_model.fit(X_train, y_train)
joblib.dump(final_model, "wine_classifier.joblib")
loaded_model = joblib.load("wine_classifier.joblib")
same_predictions = np.array_equal(
final_model.predict(X_test),
loaded_model.predict(X_test),
)
print("Loaded model test accuracy:", f"{loaded_model.score(X_test, y_test):.1%}")
print("Predictions are identical:", same_predictions)
Expected output:
Loaded model test accuracy: 100.0%
Predictions are identical: True
Only load joblib or pickle files you trust. Loading serialized Python objects can execute code.
Common errors and quick fixes
| Error / symptom | Likely cause | Fix |
|---|---|---|
NameError: name 'X_train' is not defined | You skipped the split cell | Run the data loading and train_test_split cells first |
ValueError: Found input variables with inconsistent numbers of samples | X and y lengths do not match | Print X.shape and y.shape before splitting |
| Very high train score, much lower test score | Overfitting | Reduce model complexity, use cross-validation, add data, or improve features |
| Good notebook score, bad real usage | Data leakage or mismatched preprocessing | Save and use the whole Pipeline, not only the model |
| Chart labels overlap | Figure too small or layout not adjusted | Increase figsize, rotate labels, use plt.tight_layout() |
Hands-on task
Repeat the whole workflow with load_iris():
- Print
X.shape,y.shape, feature names, and class names. - Draw a scatter plot using two features.
- Split with
train_test_split(..., stratify=y). - Train a
Pipeline(StandardScaler(), LogisticRegression(...)). - Print train/test accuracy.
- Draw a confusion matrix.
- Tune
Cwith cross-validation. - Save and reload the model with
joblib.
What should you take away from this workshop?
If Chapter 5 has one hands-on loop, it is this:
Look at the data first, split before fitting, use Pipeline for preprocessing plus model, evaluate on hidden data, improve with cross-validation, and save the complete workflow.