5.3.3 Dimensionality Reduction
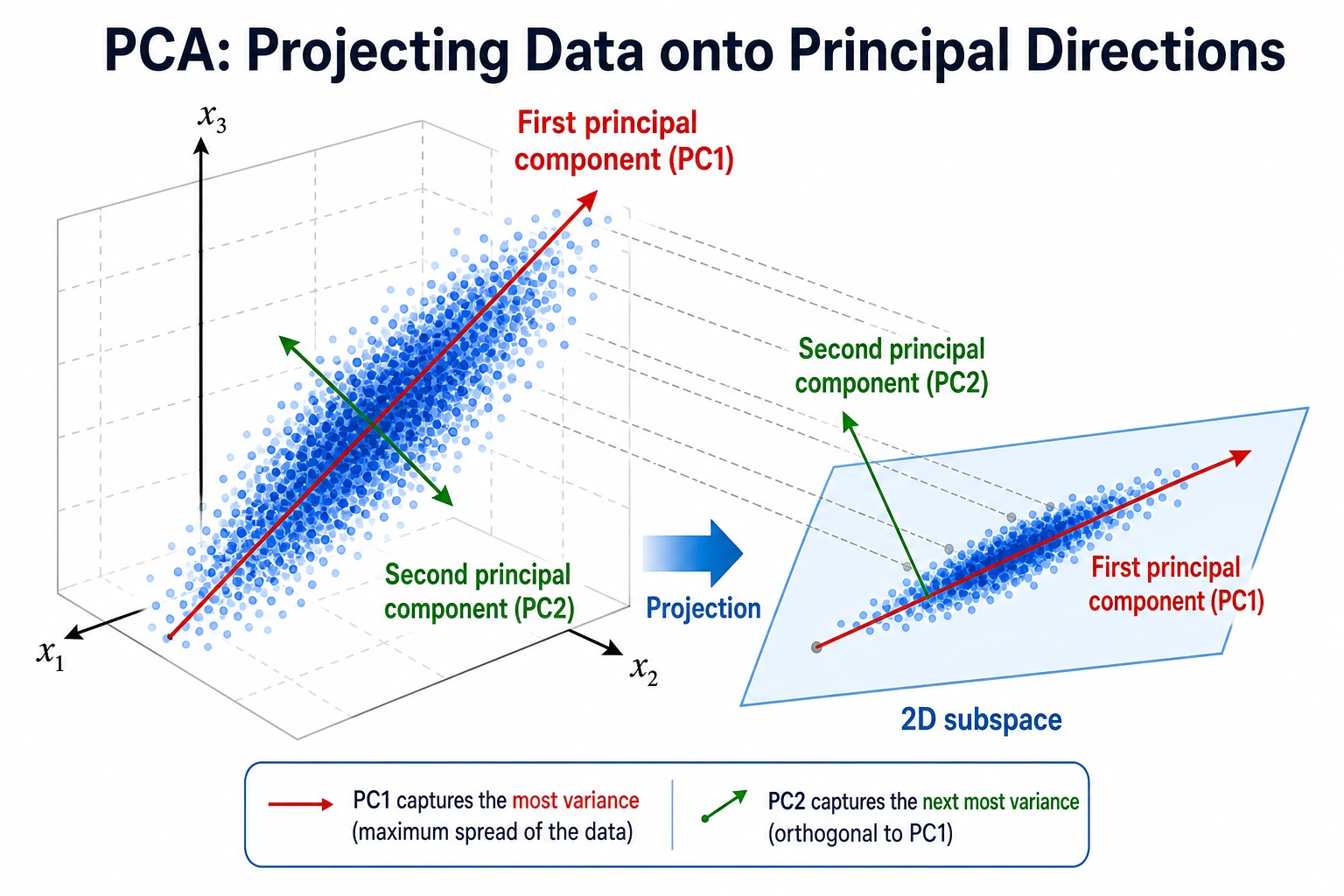
Dimensionality reduction compresses many features into fewer features. It can help with visualization, speed, noise reduction, and modeling, but each goal needs a different check.
What You Will Build
This lesson uses the handwritten digits dataset to show:
- how PCA maps high-dimensional images into 2 dimensions;
- how explained variance changes when keeping 10, 20, or 40 components;
- how PCA affects classification accuracy;
- how reconstruction error drops as more components are kept;
- when PCA, t-SNE, and UMAP should be used differently.
Look at the maps first. Dimensionality reduction is not one tool with one purpose.
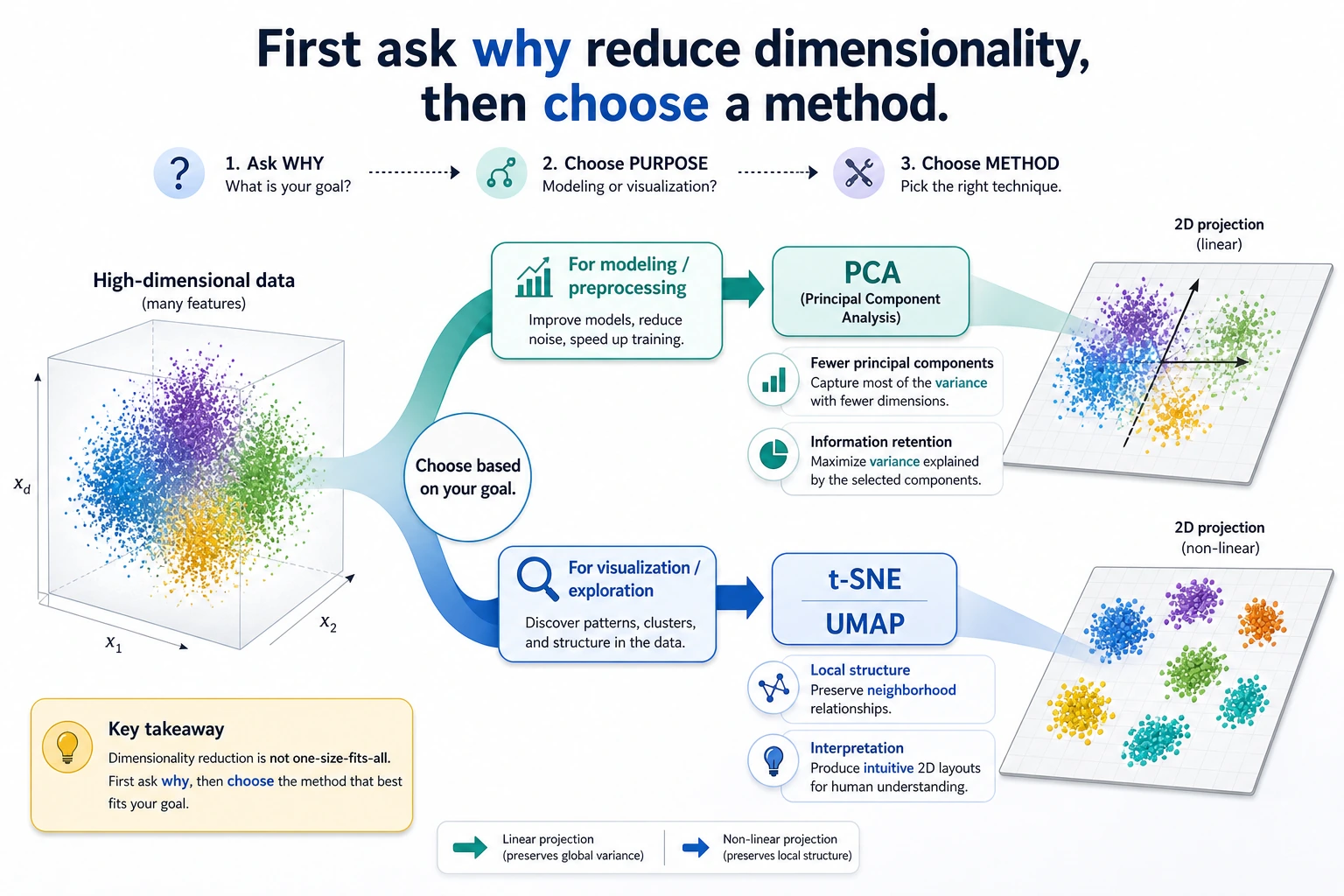
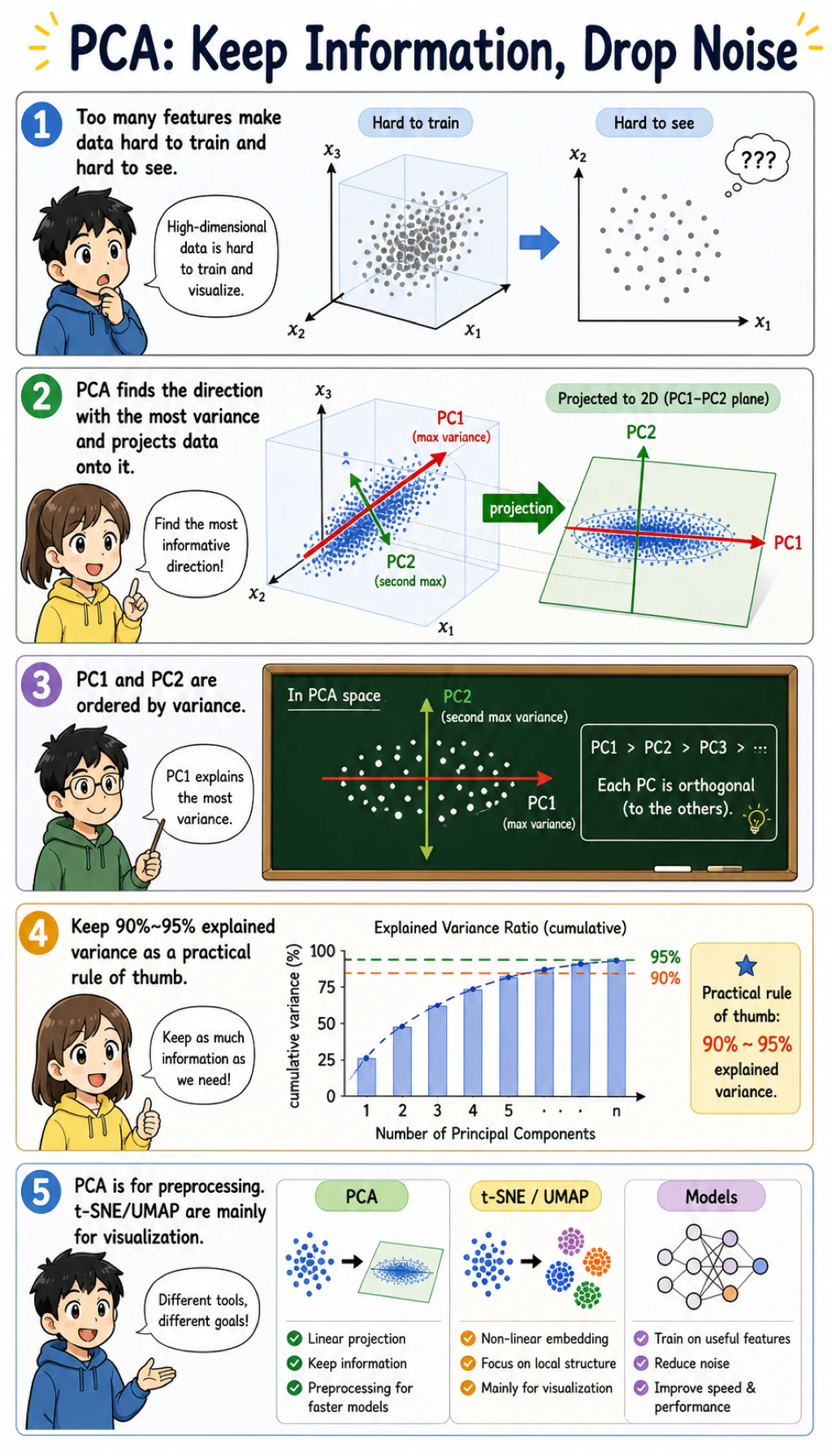
Keyword Decoder
| Term | Practical meaning |
|---|---|
dimension | One feature column, such as one pixel or one numeric field |
PCA | Principal Component Analysis; finds directions that keep as much variance as possible |
component | A new compressed feature created by PCA |
explained_variance_ratio_ | How much information-like variance each component keeps |
reconstruction | Approximate original data rebuilt from compressed components |
t-SNE | Visualization method for local neighborhood structure |
UMAP | Visualization and manifold method often used for embeddings |
Setup
python -m pip install -U scikit-learn numpy
The runnable lab uses only sklearn and NumPy. UMAP is useful in real projects, but it requires an extra package, so this beginner lab keeps the core dependency small.
Run the Complete Lab
Create pca_lab.py:
import numpy as np
from sklearn.datasets import load_digits
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
X, y = load_digits(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42, stratify=y
)
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
print("pca_2d_map")
pca2 = PCA(n_components=2, random_state=42)
X_train_2d = pca2.fit_transform(X_train_scaled)
print("shape=", X_train_2d.shape)
print("explained_variance=", np.round(pca2.explained_variance_ratio_, 3).tolist())
print("total_2d_variance=", round(float(pca2.explained_variance_ratio_.sum()), 3))
print("pca_modeling_lab")
for n in [10, 20, 40]:
model = Pipeline([
("scale", StandardScaler()),
("pca", PCA(n_components=n, random_state=42)),
("clf", LogisticRegression(max_iter=5000, random_state=42)),
])
model.fit(X_train, y_train)
pred = model.predict(X_test)
pca = model.named_steps["pca"]
print(
f"components={n:<2} "
f"variance={pca.explained_variance_ratio_.sum():.3f} "
f"accuracy={accuracy_score(y_test, pred):.3f}"
)
print("reconstruction_lab")
for n in [10, 20, 40]:
pca = PCA(n_components=n, random_state=42)
compressed = pca.fit_transform(X_train_scaled)
restored = pca.inverse_transform(compressed)
mse = mean_squared_error(X_train_scaled, restored)
print(f"components={n:<2} reconstruction_mse={mse:.3f}")
Run it:
python pca_lab.py
Expected output:
pca_2d_map
shape= (1347, 2)
explained_variance= [0.119, 0.097]
total_2d_variance= 0.216
pca_modeling_lab
components=10 variance=0.591 accuracy=0.858
components=20 variance=0.791 accuracy=0.942
components=40 variance=0.953 accuracy=0.960
reconstruction_lab
components=10 reconstruction_mse=0.390
components=20 reconstruction_mse=0.199
components=40 reconstruction_mse=0.045
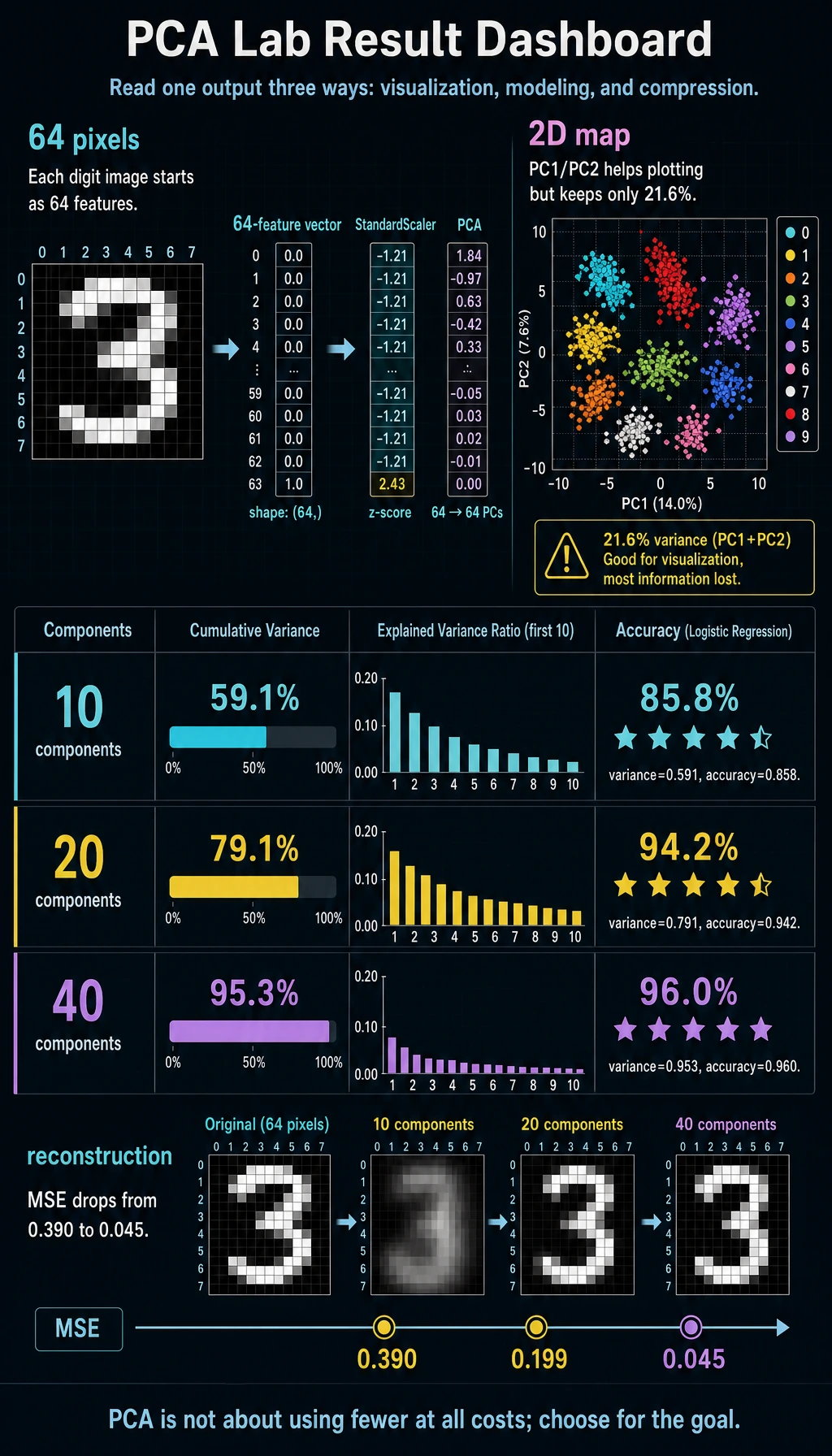

Read the 2D Result
The digits dataset has 64 pixel features. PCA with n_components=2 compresses each image into two numbers:
shape= (1347, 2)
total_2d_variance= 0.216
Two components are useful for plotting, but they keep only about 21.6% of the variance. That is fine for a quick map, but too little for a serious classifier.
Explained Variance
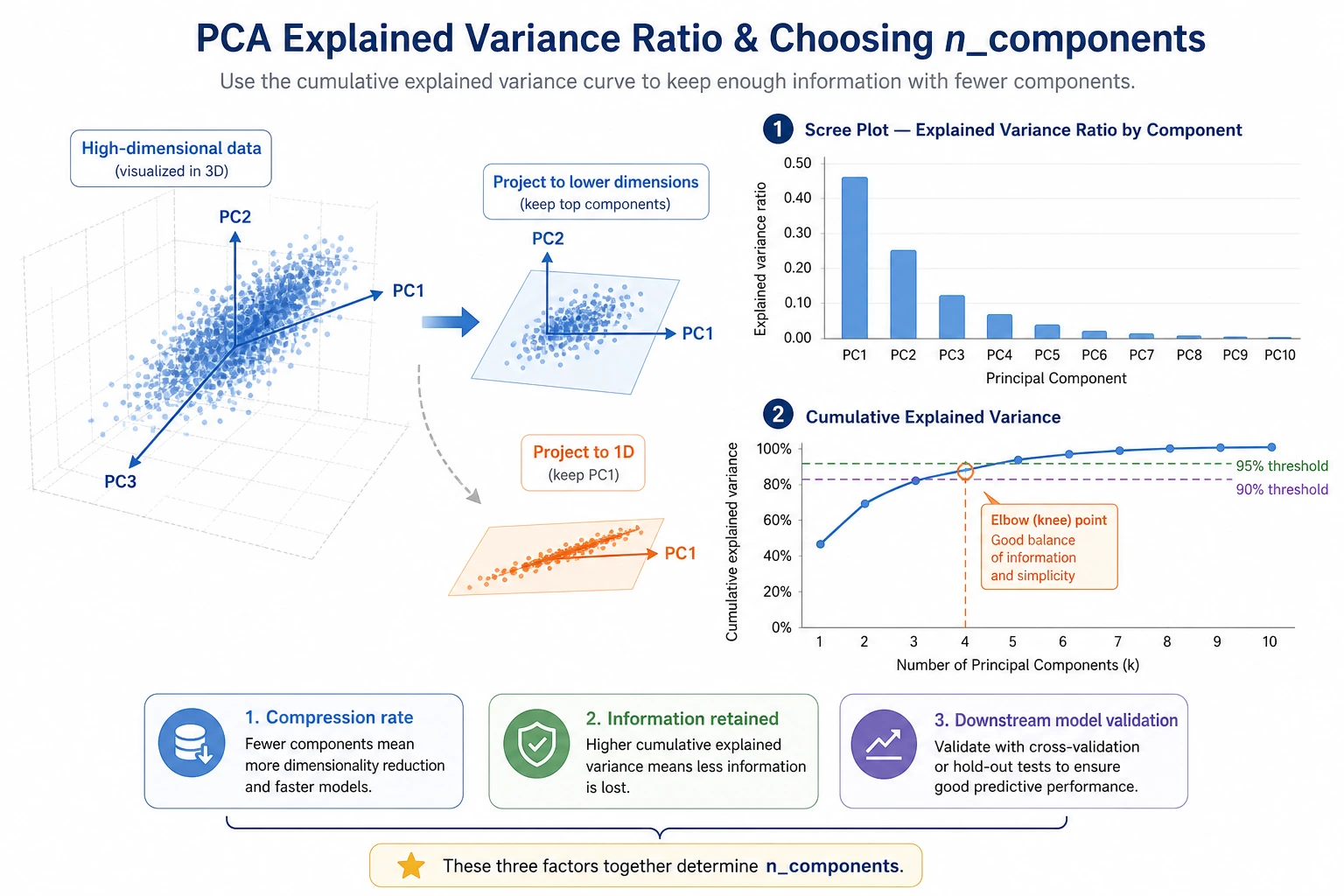
Explained variance helps you decide how much information to keep:
components=10 variance=0.591 accuracy=0.858
components=20 variance=0.791 accuracy=0.942
components=40 variance=0.953 accuracy=0.960
The useful lesson is not "always keep 95%." The useful lesson is:
- if the goal is visualization,
2or3components may be enough; - if the goal is modeling, compare accuracy or the metric your project uses;
- if the goal is compression, compare reconstruction error and storage cost.
Reconstruction Error
Reconstruction asks: after compression, how much of the original data can be rebuilt?
components=10 reconstruction_mse=0.390
components=40 reconstruction_mse=0.045
More components make reconstruction better, but they also keep more dimensions. The right number is a trade-off between compactness and useful information.
PCA in a Model Pipeline
The modeling block uses:
Pipeline([
("scale", StandardScaler()),
("pca", PCA(n_components=n, random_state=42)),
("clf", LogisticRegression(max_iter=5000, random_state=42)),
])
This order matters:
- split train/test first;
- fit scaling on training data only;
- fit PCA on training data only;
- train the model on compressed training features;
- evaluate on transformed test features.
Putting scaling and PCA in a pipeline helps prevent data leakage during cross-validation.
PCA, t-SNE, and UMAP
| Method | Best use | Important warning |
|---|---|---|
| PCA | compression, preprocessing, fast 2D overview | linear method; may miss curved structure |
| t-SNE | local-neighborhood visualization | distances between far clusters can be misleading |
| UMAP | embedding visualization and neighborhood exploration | needs extra package; tune parameters and verify stability |
For beginners, the safest order is:
- Start with PCA because it is fast and interpretable.
- Use t-SNE or UMAP for visualization, not as your first production feature pipeline.
- If dimensionality reduction changes a model, validate with cross-validation.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| PCA result dominated by one feature | features are not scaled | use StandardScaler before PCA |
| 2D plot looks nice but model is weak | 2D kept too little variance | use more components for modeling |
| Accuracy drops sharply after PCA | too many useful features were discarded | increase n_components, compare with no-PCA baseline |
| Cross-validation score is too good | PCA fitted before split | put PCA inside Pipeline |
| t-SNE/UMAP plot looks overinterpreted | visualization layout is not a proof | check stability and downstream usefulness |
Practice
- Change PCA components to
[5, 15, 30, 50]. Where does accuracy stop improving? - Run the classifier without PCA. Is PCA helping speed, accuracy, or only compression?
- Remove
StandardScaler. How does explained variance change? - Use
PCA(n_components=0.95)and print the number of components selected. - Use the 2D PCA output to draw a scatter plot colored by digit label.
Pass Check
You are done when you can explain:
- PCA creates new compressed features called components;
- 2D PCA is useful for visualization but may discard too much information for modeling;
- explained variance is a guide, not an automatic target;
- PCA must be fitted inside the training pipeline;
- t-SNE and UMAP are mainly visualization tools unless you validate them carefully.