5.3.2 Clustering Algorithms
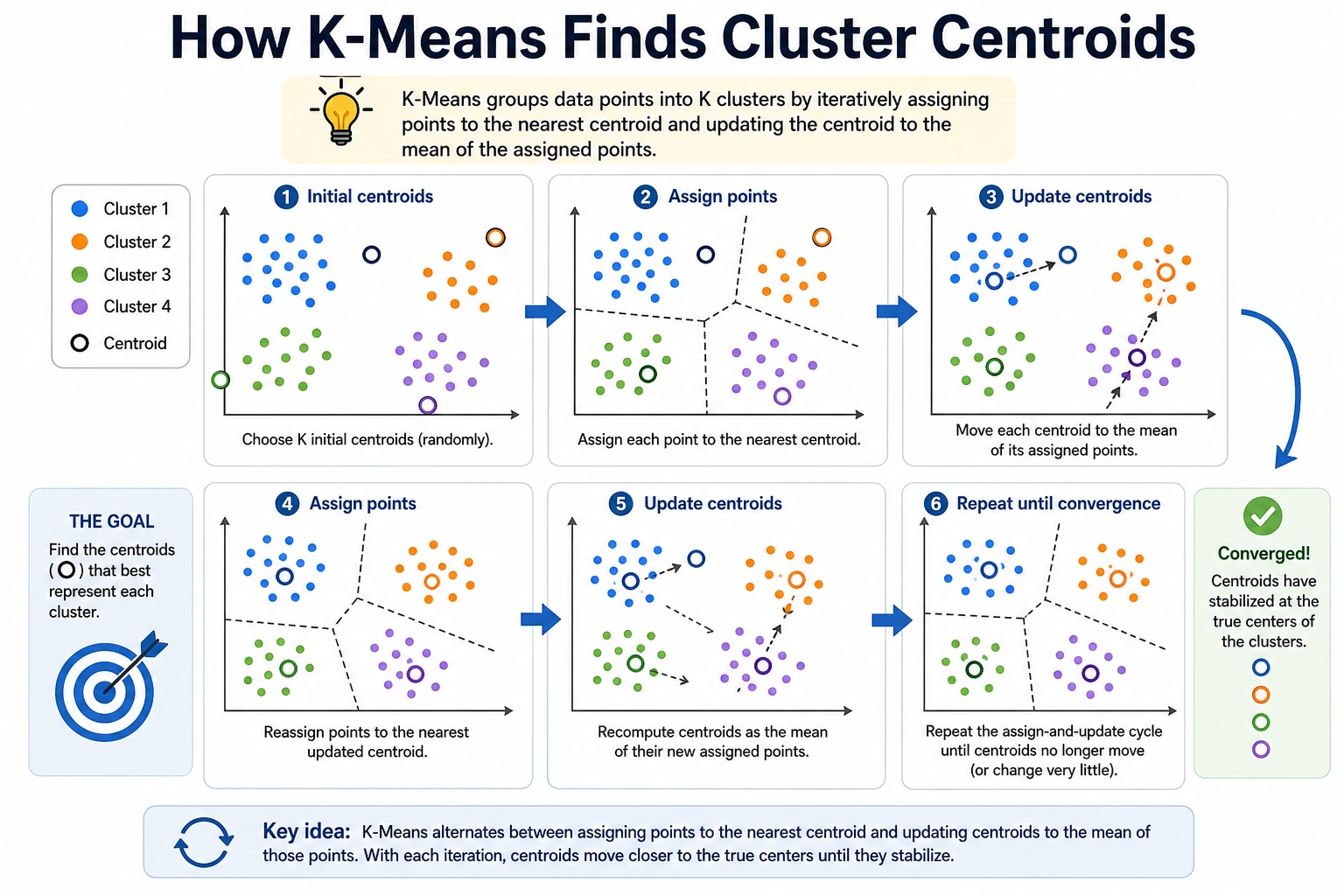
Clustering means grouping similar samples without labels. The result is not the one true answer; it is a hypothesis about structure that must be checked with metrics, plots, and domain meaning.
What You Will Build
This lesson gives you one practical clustering lab:
- choose
Kfor K-Means with inertia and silhouette score; - inspect K-Means cluster centers;
- compare K-Means with DBSCAN on curved data;
- tune DBSCAN's
eps; - run hierarchical clustering as an inspection-friendly alternative.
Read the maps first. Clustering is mostly about matching the algorithm's assumption to the data shape.
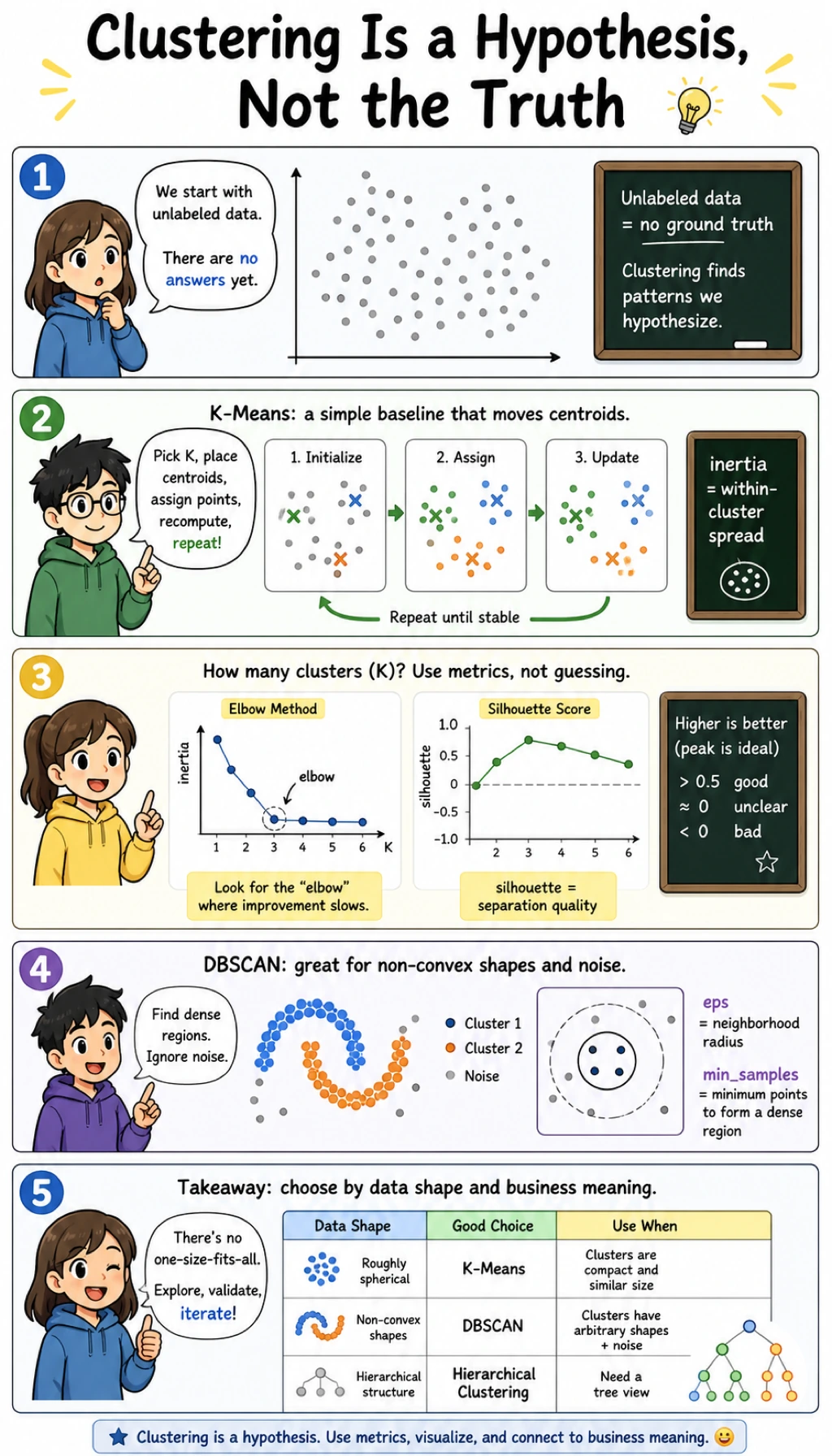
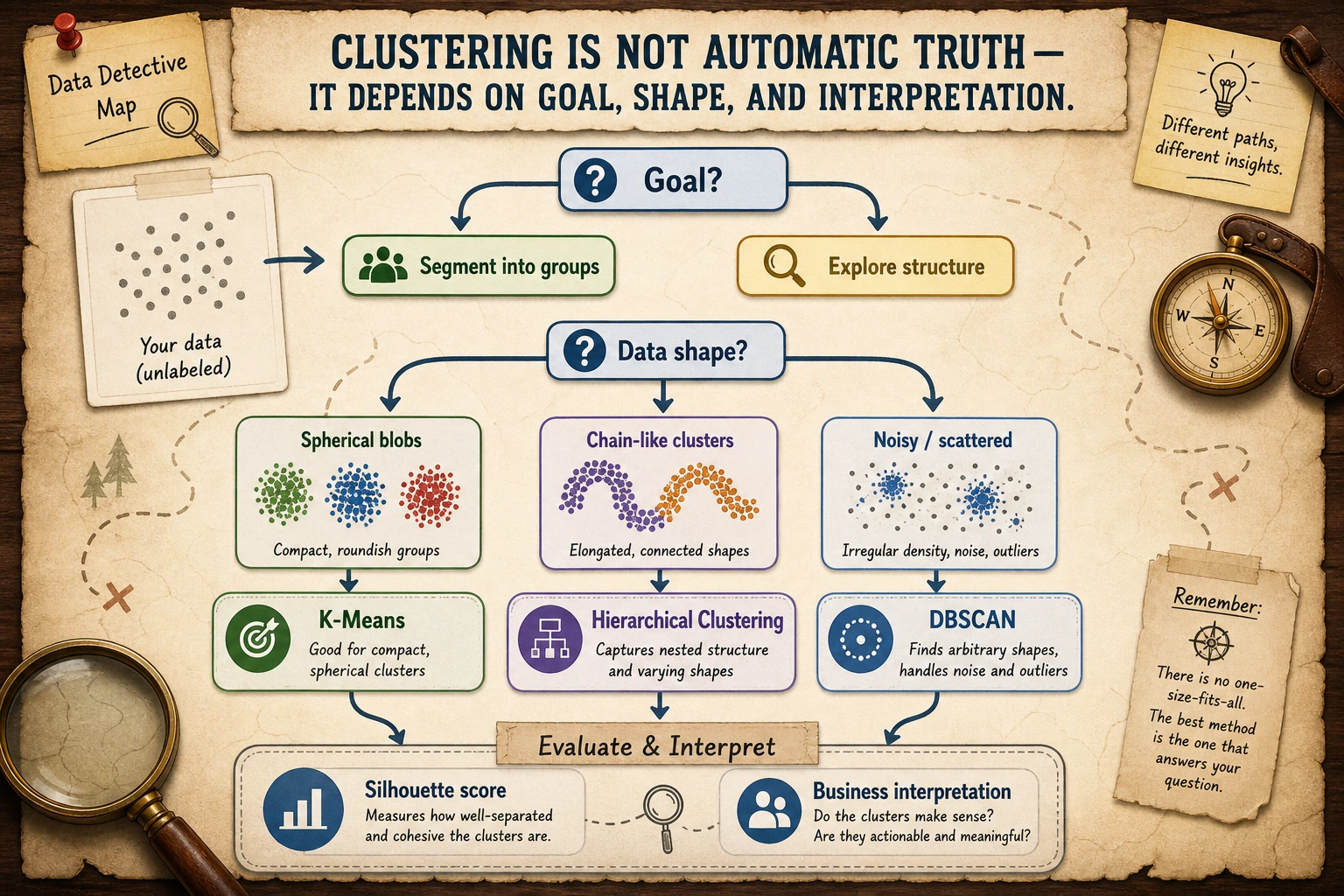

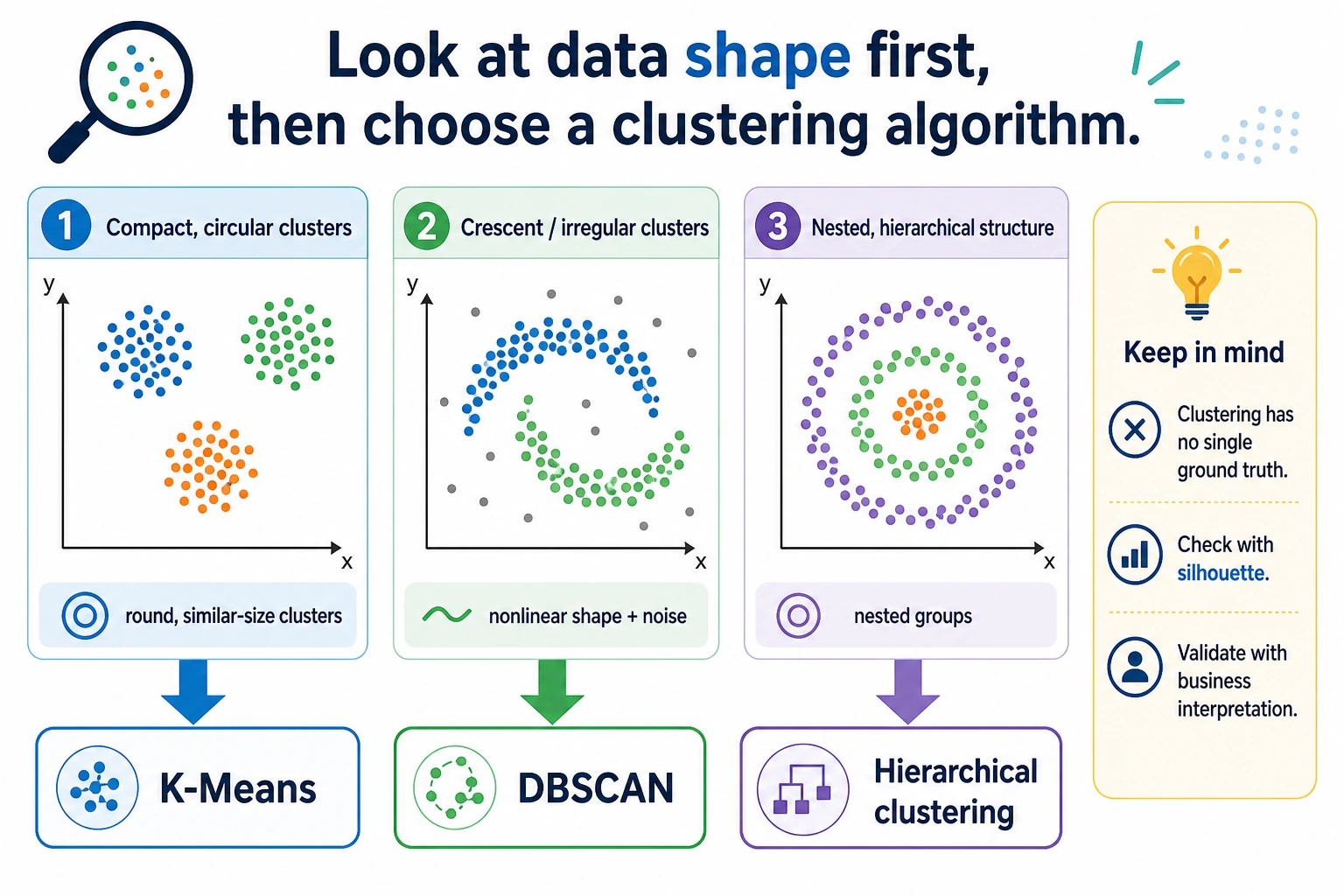
Keyword Decoder
| Term | Practical meaning |
|---|---|
cluster | A group of points that look similar under the chosen features |
centroid | The center of a K-Means cluster |
inertia_ | Within-cluster squared distance; lower is more compact but always drops as K grows |
silhouette_score | Measures both compactness and separation; higher is usually better |
eps | DBSCAN neighborhood radius |
min_samples | Minimum neighbors needed for a dense DBSCAN core point |
noise | DBSCAN label -1, meaning "not assigned to a dense cluster" |
linkage | Hierarchical clustering rule for merging groups |
Setup
python -m pip install -U scikit-learn numpy
All examples scale features first. Clustering is usually distance-based, so feature scale changes the meaning of "similar."
Run the Complete Lab
Create clustering_lab.py:
import numpy as np
from sklearn.cluster import AgglomerativeClustering, DBSCAN, KMeans
from sklearn.datasets import make_blobs, make_moons
from sklearn.metrics import adjusted_rand_score, silhouette_score
from sklearn.preprocessing import StandardScaler
# Round blob clusters: good K-Means demo.
X_blob, y_blob = make_blobs(n_samples=360, centers=3, cluster_std=0.85, random_state=42)
X_blob = StandardScaler().fit_transform(X_blob)
print("kmeans_k_selection")
for k in [2, 3, 4, 5]:
model = KMeans(n_clusters=k, n_init="auto", random_state=42)
labels = model.fit_predict(X_blob)
print(
f"k={k} inertia={model.inertia_:6.1f} "
f"silhouette={silhouette_score(X_blob, labels):.3f}"
)
best = KMeans(n_clusters=3, n_init="auto", random_state=42)
labels = best.fit_predict(X_blob)
print("kmeans_centers")
print(np.round(best.cluster_centers_, 2))
print("kmeans_ari=", round(adjusted_rand_score(y_blob, labels), 3))
# Curved clusters: DBSCAN is a better fit than K-Means.
X_moon, y_moon = make_moons(n_samples=400, noise=0.08, random_state=42)
X_moon = StandardScaler().fit_transform(X_moon)
print("shape_mismatch_lab")
kmeans = KMeans(n_clusters=2, n_init="auto", random_state=42)
km_labels = kmeans.fit_predict(X_moon)
print("kmeans_moon_ari=", round(adjusted_rand_score(y_moon, km_labels), 3))
for eps in [0.15, 0.25, 0.35]:
db = DBSCAN(eps=eps, min_samples=5)
db_labels = db.fit_predict(X_moon)
clusters = len(set(db_labels)) - (1 if -1 in db_labels else 0)
noise = int(np.sum(db_labels == -1))
print(
f"dbscan eps={eps:.2f} clusters={clusters} noise={noise} "
f"ari={adjusted_rand_score(y_moon, db_labels):.3f}"
)
print("hierarchical_lab")
agg = AgglomerativeClustering(n_clusters=3, linkage="ward")
agg_labels = agg.fit_predict(X_blob)
print("agglomerative_ari=", round(adjusted_rand_score(y_blob, agg_labels), 3))
Run it:
python clustering_lab.py
Expected output:
kmeans_k_selection
k=2 inertia= 417.4 silhouette=0.527
k=3 inertia= 16.4 silhouette=0.869
k=4 inertia= 14.6 silhouette=0.690
k=5 inertia= 11.9 silhouette=0.532
kmeans_centers
[[-0.2 1.17]
[-1.09 -1.25]
[ 1.29 0.08]]
kmeans_ari= 1.0
shape_mismatch_lab
kmeans_moon_ari= 0.475
dbscan eps=0.15 clusters=12 noise=37 ari=0.312
dbscan eps=0.25 clusters=2 noise=1 ari=0.995
dbscan eps=0.35 clusters=2 noise=1 ari=0.995
hierarchical_lab
agglomerative_ari= 1.0
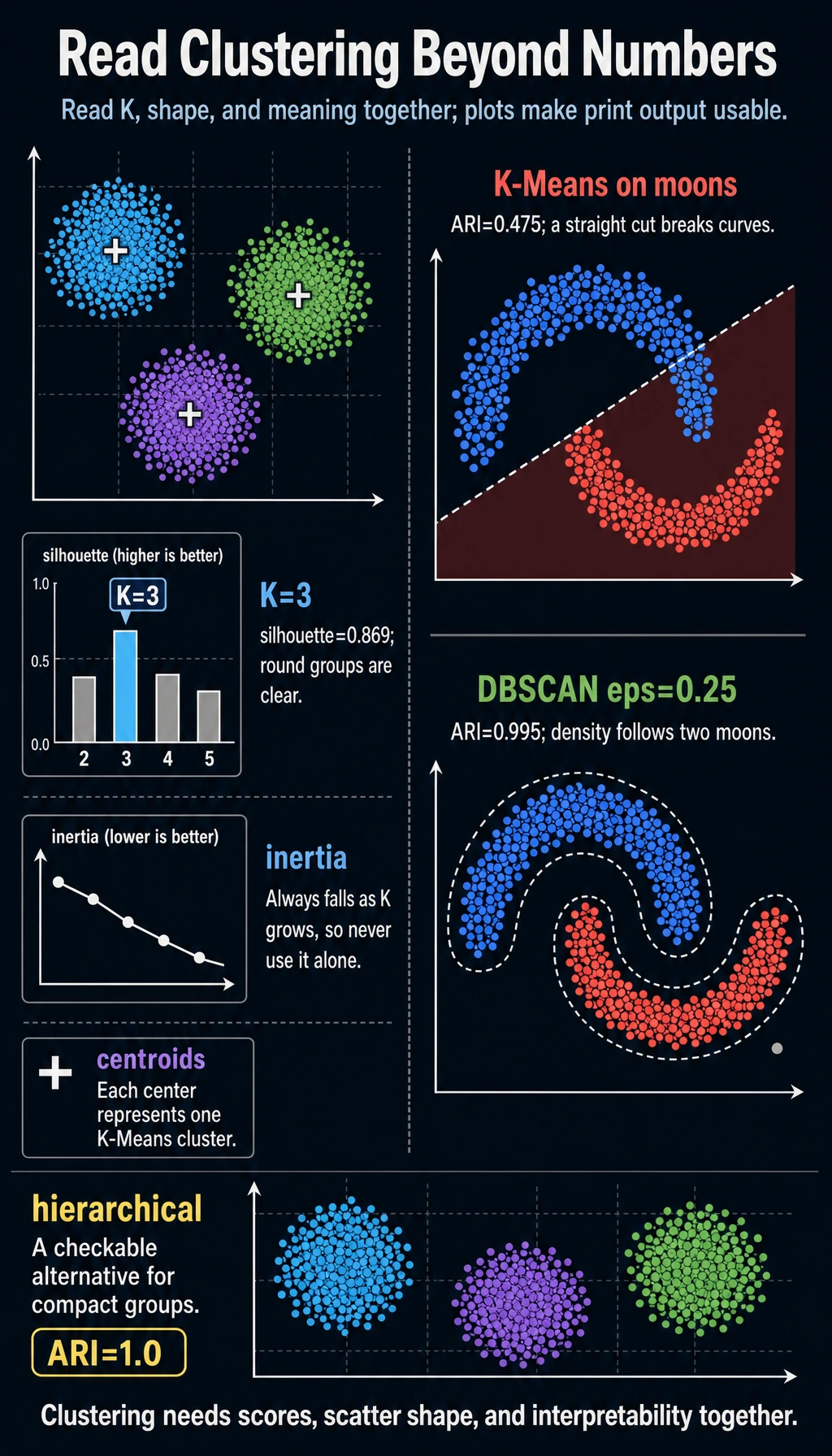
adjusted_rand_score uses the hidden synthetic labels only so this teaching lab can verify behavior. In real clustering work, you usually do not have labels, so you rely on metrics, visualization, and business interpretation.
K-Means: Choose K
K-Means repeats three steps:
- place
Kcentroids; - assign each point to the nearest centroid;
- move each centroid to the mean of its assigned points.
The lab compares candidate K values:
k=2 inertia= 417.4 silhouette=0.527
k=3 inertia= 16.4 silhouette=0.869
k=4 inertia= 14.6 silhouette=0.690
Here K=3 is the best practical choice:
- inertia drops sharply from
K=2toK=3; - silhouette is highest at
K=3; - adding more clusters lowers inertia but makes the grouping less separated.
Do not choose K from inertia alone. Inertia always improves when K increases, because smaller groups are easier to fit.
K-Means Assumption
K-Means works best when clusters are:
- roughly round;
- similarly sized;
- separated by distance;
- measured on comparable feature scales.
It struggles when clusters are curved, nested, noisy, or very different in density.
DBSCAN: Find Dense Regions
DBSCAN does not ask for K. It asks:
Which points have enough neighbors inside radius
eps?
That makes it useful for curved shapes and noisy data. The lab shows the shape mismatch:
kmeans_moon_ari= 0.475
dbscan eps=0.25 clusters=2 noise=1 ari=0.995
K-Means tries to cut the moons into distance-based regions. DBSCAN follows dense curves, so it recovers the two moon shapes.
The key parameter is eps:
dbscan eps=0.15 clusters=12 noise=37
dbscan eps=0.25 clusters=2 noise=1
If eps is too small, DBSCAN breaks one real group into many small pieces. If eps is too large, it can merge groups together.
Hierarchical Clustering
Hierarchical clustering repeatedly merges nearby groups. It is useful when you want to inspect nested relationships or create a dendrogram outside this minimal script.
In the lab:
agglomerative_ari= 1.0
linkage="ward" works well on the round blob data because it prefers compact clusters. For non-round shapes, it may not be enough by itself.
Algorithm Choice
| Data shape / goal | Good first choice | Why |
|---|---|---|
| Round, compact groups | K-Means | fast, simple, strong baseline |
Unknown K, noisy curved shapes | DBSCAN | can mark noise and follow dense regions |
| Need hierarchy inspection | Agglomerative clustering | shows merge structure |
| Very high-dimensional embeddings | K-Means or HDBSCAN-style tools | compare with visualization and retrieval checks |
| Business segmentation | K-Means baseline plus domain review | groups must be actionable, not only pretty |
For experienced readers: clustering should be evaluated as a workflow, not just an algorithm score. Check stability under resampling, feature changes, scaling choices, and different random seeds.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| K-Means result changes a lot | initialization instability | use n_init="auto", try several seeds |
More K always looks better by inertia | inertia always decreases with K | also use silhouette and business interpretability |
| DBSCAN returns mostly noise | eps too small, features not scaled | scale features, increase eps |
| DBSCAN returns one giant cluster | eps too large | decrease eps |
| Clusters look nice but are useless | features do not match actions | define what each cluster will change in the product |
Practice
- Change
cluster_stdinmake_blobs()from0.85to1.5. How does silhouette change? - Add
K=6to the K-Means loop. Does inertia improve? Does silhouette improve? - Try
min_samples=10in DBSCAN. What happens to noise count? - Replace the synthetic data with customer data. Scale numeric features first, then explain each cluster in plain language.
- Run the same clustering twice with different seeds. Are the groups stable enough to trust?
Pass Check
You are done when you can explain:
- clustering creates a hypothesis, not a guaranteed truth;
- K-Means is a strong baseline for round, compact groups;
- inertia alone cannot choose
K; - DBSCAN is useful for dense curved shapes and noise;
- the final cluster names must be validated by real-world meaning.