5.4.3 Cross-Validation
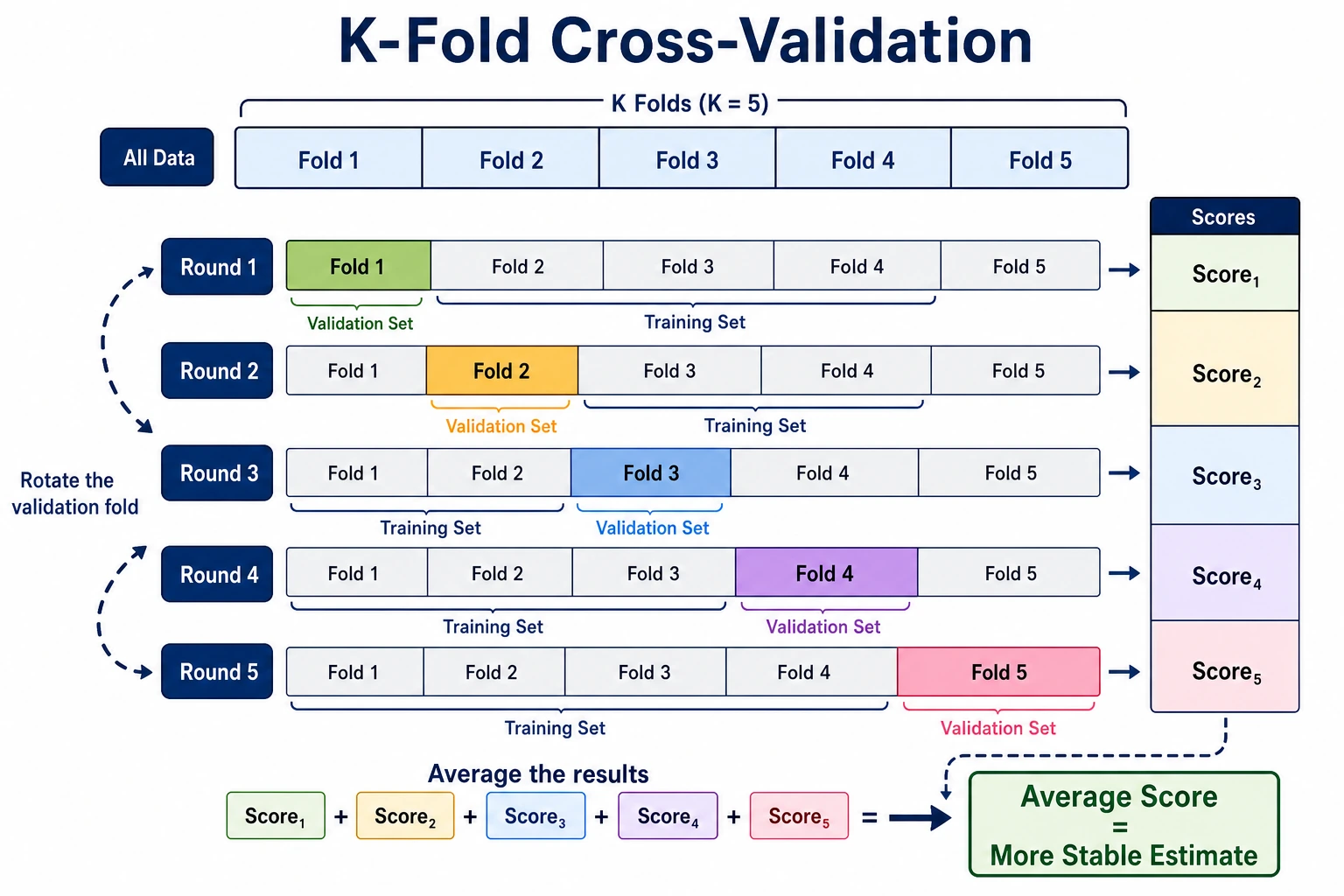
A single train-test split is a snapshot. Cross-validation gives you a more stable estimate by testing the model across several different validation folds.
What You Will Build
This lesson shows:
- why one train-test split can be noisy;
- how to use
StratifiedKFoldfor classification; - how to evaluate several metrics with
cross_validate; - why preprocessing must stay inside
Pipeline; - when random K-Fold is wrong, especially for time series.
Setup
python -m pip install -U scikit-learn numpy
Run the Complete Lab
Create cv_lab.py:
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.model_selection import StratifiedKFold, cross_validate, train_test_split
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
X, y = load_breast_cancer(return_X_y=True)
print("single_split_variance")
for seed in [1, 2, 3, 4, 5]:
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=seed, stratify=y
)
model = Pipeline([
("scale", StandardScaler()),
("clf", LogisticRegression(max_iter=2000, random_state=42)),
])
model.fit(X_train, y_train)
print(f"seed={seed} accuracy={accuracy_score(y_test, model.predict(X_test)):.3f}")
print("cross_validation_lab")
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
model = Pipeline([
("scale", StandardScaler()),
("clf", LogisticRegression(max_iter=2000, random_state=42)),
])
result = cross_validate(
model,
X,
y,
cv=cv,
scoring=["accuracy", "precision", "recall", "f1"],
)
for i, score in enumerate(result["test_accuracy"], start=1):
print(f"fold={i} accuracy={score:.3f}")
print(
"summary "
f"accuracy={np.mean(result['test_accuracy']):.3f}+/-{np.std(result['test_accuracy']):.3f} "
f"precision={np.mean(result['test_precision']):.3f} "
f"recall={np.mean(result['test_recall']):.3f} "
f"f1={np.mean(result['test_f1']):.3f}"
)
Run it:
python cv_lab.py
Expected output:
single_split_variance
seed=1 accuracy=0.965
seed=2 accuracy=0.972
seed=3 accuracy=0.986
seed=4 accuracy=0.972
seed=5 accuracy=0.979
cross_validation_lab
fold=1 accuracy=0.974
fold=2 accuracy=0.947
fold=3 accuracy=0.965
fold=4 accuracy=0.991
fold=5 accuracy=0.991
summary accuracy=0.974+/-0.017 precision=0.968 recall=0.992 f1=0.979
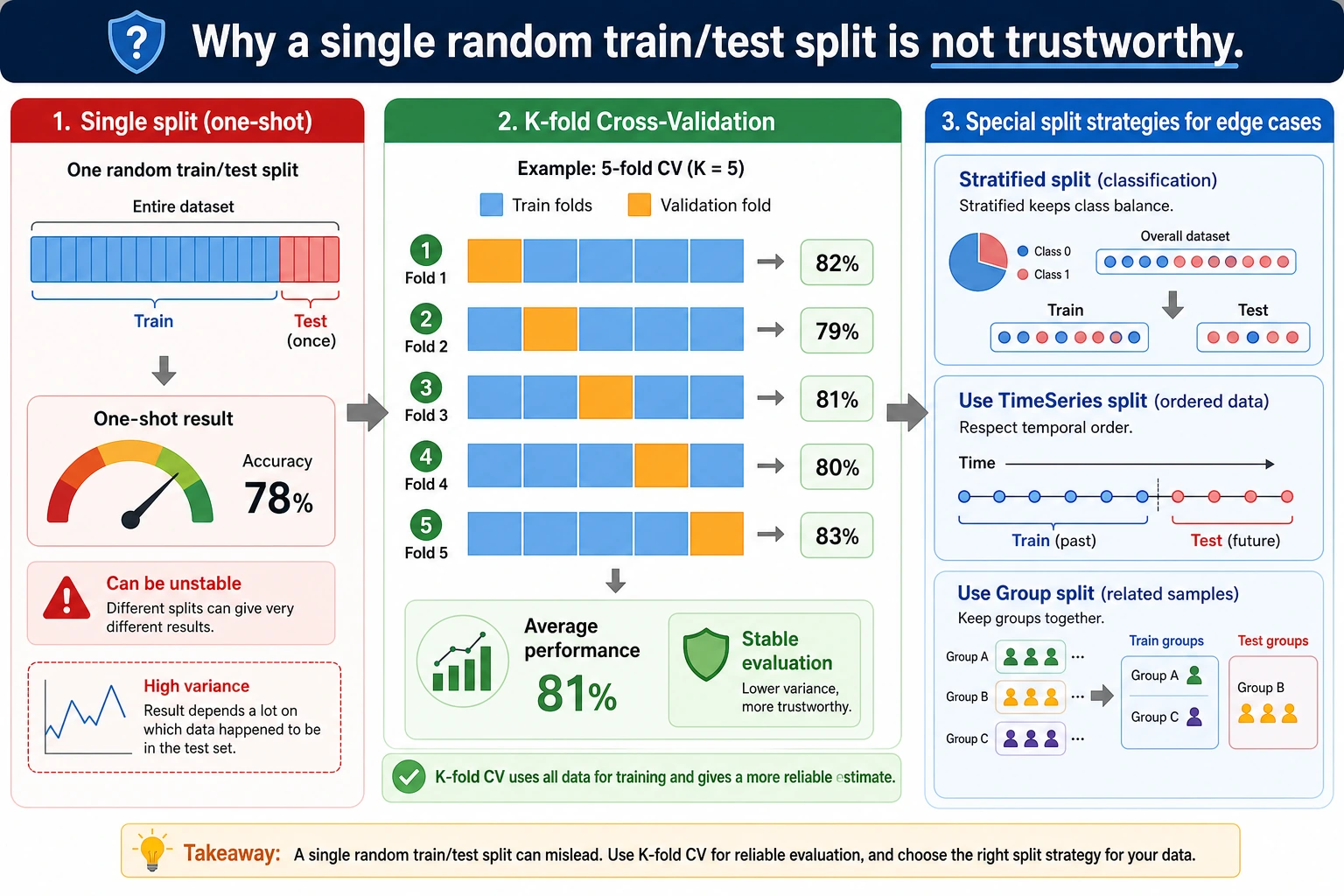
Why One Split Is Not Enough
The same model gets different scores with different random splits:
seed=1 accuracy=0.965
seed=3 accuracy=0.986
Neither number is fake. They are just different snapshots. Cross-validation asks: "Across several snapshots, what is the average performance and how much does it vary?"
Stratified K-Fold
For classification, use StratifiedKFold first. It keeps the class ratio similar in each fold, which is especially important for imbalanced datasets.
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
Use K=5 as a practical default:
- less noisy than one split;
- cheaper than 10-fold on large data;
- easy to explain to teammates.
Use a Leakage-Safe Pipeline
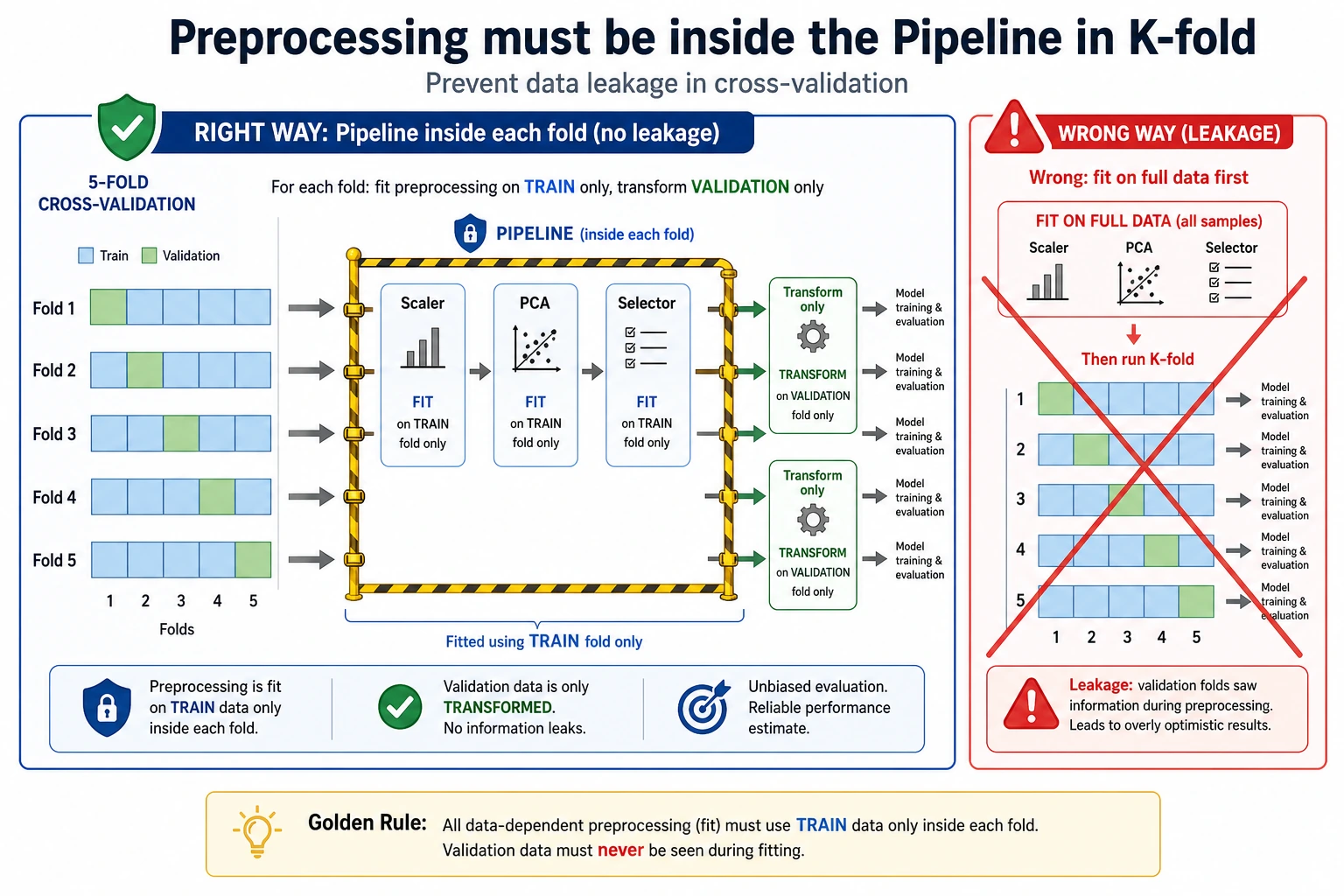
This is the safe pattern:
Pipeline([
("scale", StandardScaler()),
("clf", LogisticRegression(max_iter=2000, random_state=42)),
])
During cross-validation, each fold must fit its scaler only on that fold's training portion. If you scale all data before CV, information from validation folds leaks into training.
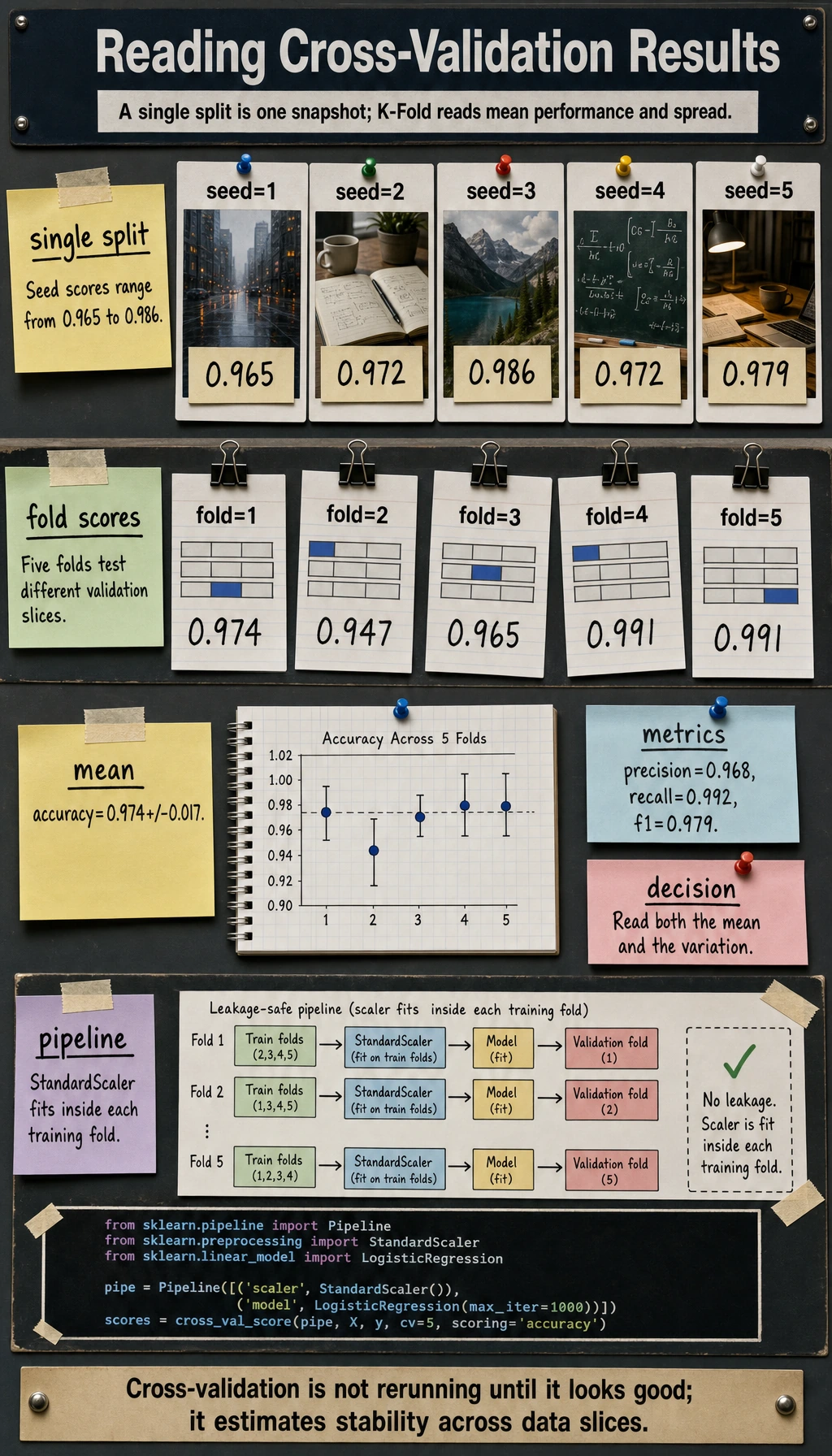
Read Mean and Variance
The summary is more useful than one fold:
summary accuracy=0.974+/-0.017 precision=0.968 recall=0.992 f1=0.979
Read it as:
- average accuracy is about
0.974; - fold-to-fold variation is about
0.017; - recall is very high, which matters if missing positives is costly.
If standard deviation is large, the model may be unstable, the dataset may be small, or some folds may contain harder cases.
When K-Fold Is Wrong
Do not shuffle randomly when:
- the data is time series;
- rows from the same user/session/device can appear in both train and validation;
- examples are grouped by patient, customer, document, or experiment;
- future information would leak into the past.
Use a split that matches the real deployment situation: TimeSeriesSplit, group splits, or a chronological holdout.
Practical Choice Guide
| Situation | Use |
|---|---|
| Basic classification | StratifiedKFold(n_splits=5, shuffle=True) |
| Regression | KFold(n_splits=5, shuffle=True) |
| Time series | TimeSeriesSplit or chronological validation |
| Same entity appears many times | group-aware splitting |
| Hyperparameter tuning | nested CV or a final untouched test set |
For experienced readers: after model selection, keep one final holdout set or production-like backtest that was not used during tuning.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| CV score much higher than test score | leakage or over-tuning | put preprocessing in pipeline; keep final holdout |
| Fold scores vary wildly | small data or hard segments | inspect fold composition and segment metrics |
| Classification fold has no positives | non-stratified split | use StratifiedKFold |
| Time-series model looks too good | future data leaked | validate chronologically |
| CV takes too long | too many folds or heavy model | use fewer folds or faster baseline first |
Practice
- Change
n_splitsto3and10. How do mean and standard deviation change? - Remove
stratify=yfrom the single split. Does the score become less stable? - Add
roc_aucto the scoring list. - Move
StandardScaler()outside the pipeline intentionally, then explain why that is unsafe. - Design a validation split for user events where each user has many rows.
Pass Check
You are done when you can explain:
- one train-test split is only one snapshot;
- K-Fold estimates average performance and variability;
- classification should usually use stratified folds;
- preprocessing must be inside the pipeline;
- validation strategy must match deployment data flow.