5.4.5 Hyperparameter Tuning
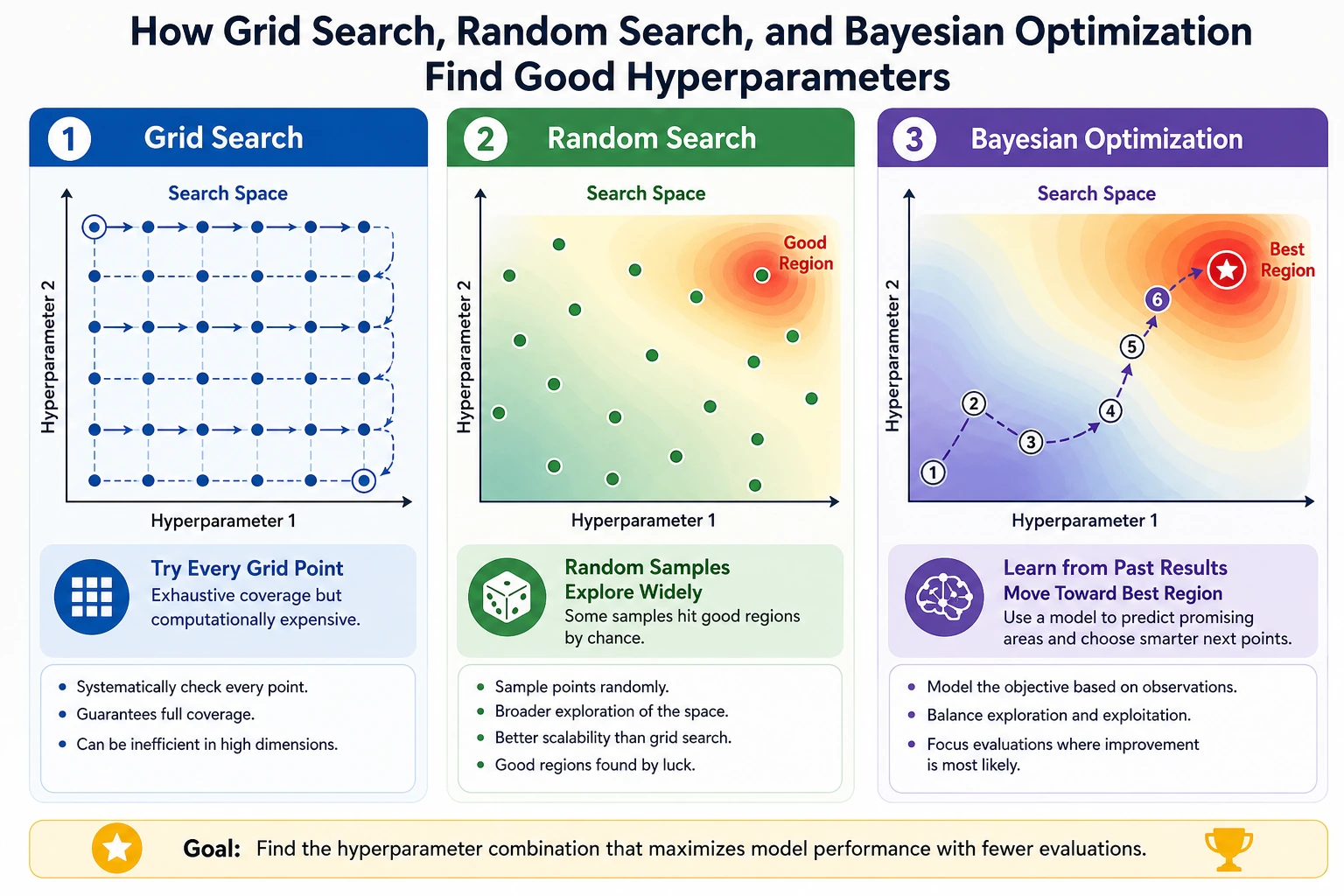
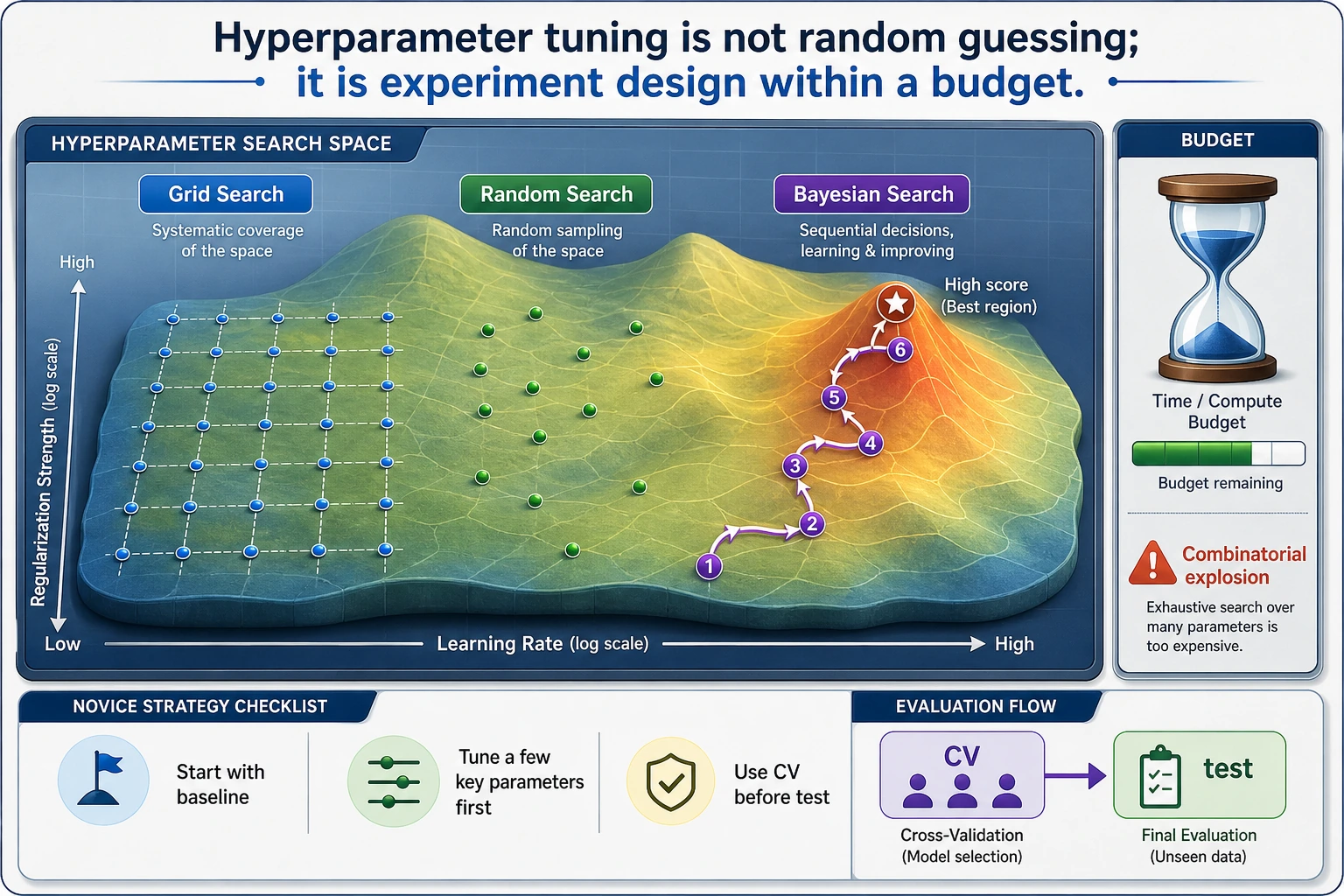
Hyperparameter tuning is not "trying settings until the test score looks good." A safe tuning workflow searches on training folds, chooses by cross-validation, and checks once on a final holdout.
What You Will Build
This lesson shows:
- the difference between parameters and hyperparameters;
- how to use
GridSearchCV; - how to use
RandomizedSearchCVwhen the search space grows; - how to keep a final holdout untouched;
- how to avoid over-tuning.
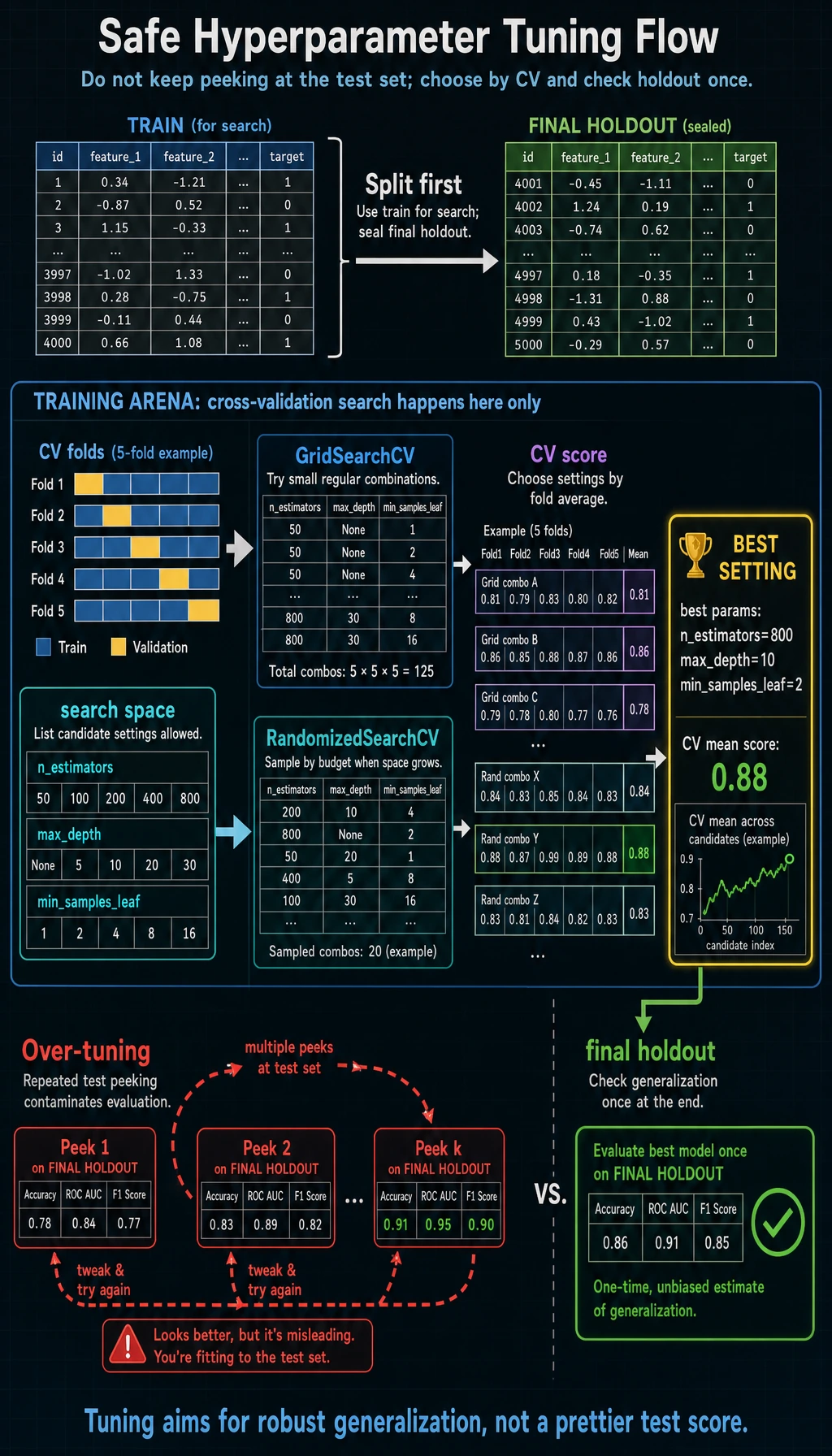

Keyword Decoder
| Term | Practical meaning |
|---|---|
| parameter | learned by the model during fit() |
| hyperparameter | chosen by you before training, such as tree depth |
| search space | the candidate values you allow the search to try |
| CV score | cross-validation score used to choose settings |
| final holdout | untouched data used once after tuning |
| budget | number of combinations or trials you can afford |
Setup
python -m pip install -U scikit-learn
Run the Complete Lab
Create tuning_lab.py:
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, f1_score, recall_score
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV, StratifiedKFold, train_test_split
X, y = load_breast_cancer(return_X_y=True)
X_train, X_final, y_train, y_final = train_test_split(
X, y, test_size=0.2, random_state=42, stratify=y
)
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
print("grid_search_lab")
grid = GridSearchCV(
RandomForestClassifier(random_state=42),
param_grid={
"n_estimators": [80, 160],
"max_depth": [3, 5, None],
"min_samples_leaf": [1, 3],
},
scoring="f1",
cv=cv,
n_jobs=-1,
)
grid.fit(X_train, y_train)
print("best_params=", grid.best_params_)
print(f"best_cv_f1={grid.best_score_:.3f}")
final_pred = grid.best_estimator_.predict(X_final)
print(
f"final accuracy={accuracy_score(y_final, final_pred):.3f} "
f"recall={recall_score(y_final, final_pred):.3f} "
f"f1={f1_score(y_final, final_pred):.3f}"
)
print("random_search_lab")
random_search = RandomizedSearchCV(
RandomForestClassifier(random_state=42),
param_distributions={
"n_estimators": [60, 100, 160, 220],
"max_depth": [3, 5, 8, None],
"min_samples_leaf": [1, 2, 3, 5],
"max_features": ["sqrt", "log2", None],
},
n_iter=8,
scoring="f1",
cv=cv,
random_state=42,
n_jobs=-1,
)
random_search.fit(X_train, y_train)
print("best_params=", random_search.best_params_)
print(f"best_cv_f1={random_search.best_score_:.3f}")
print("top_3_grid_results")
rows = sorted(
zip(grid.cv_results_["mean_test_score"], grid.cv_results_["params"]),
key=lambda item: item[0],
reverse=True,
)[:3]
for score, params in rows:
print(f"score={score:.3f} params={params}")
Run it:
python tuning_lab.py
Expected output:
grid_search_lab
best_params= {'max_depth': 5, 'min_samples_leaf': 3, 'n_estimators': 80}
best_cv_f1=0.968
final accuracy=0.956 recall=0.972 f1=0.966
random_search_lab
best_params= {'n_estimators': 100, 'min_samples_leaf': 1, 'max_features': 'log2', 'max_depth': 8}
best_cv_f1=0.972
top_3_grid_results
score=0.968 params={'max_depth': 5, 'min_samples_leaf': 3, 'n_estimators': 80}
score=0.968 params={'max_depth': 5, 'min_samples_leaf': 3, 'n_estimators': 160}
score=0.968 params={'max_depth': None, 'min_samples_leaf': 3, 'n_estimators': 160}
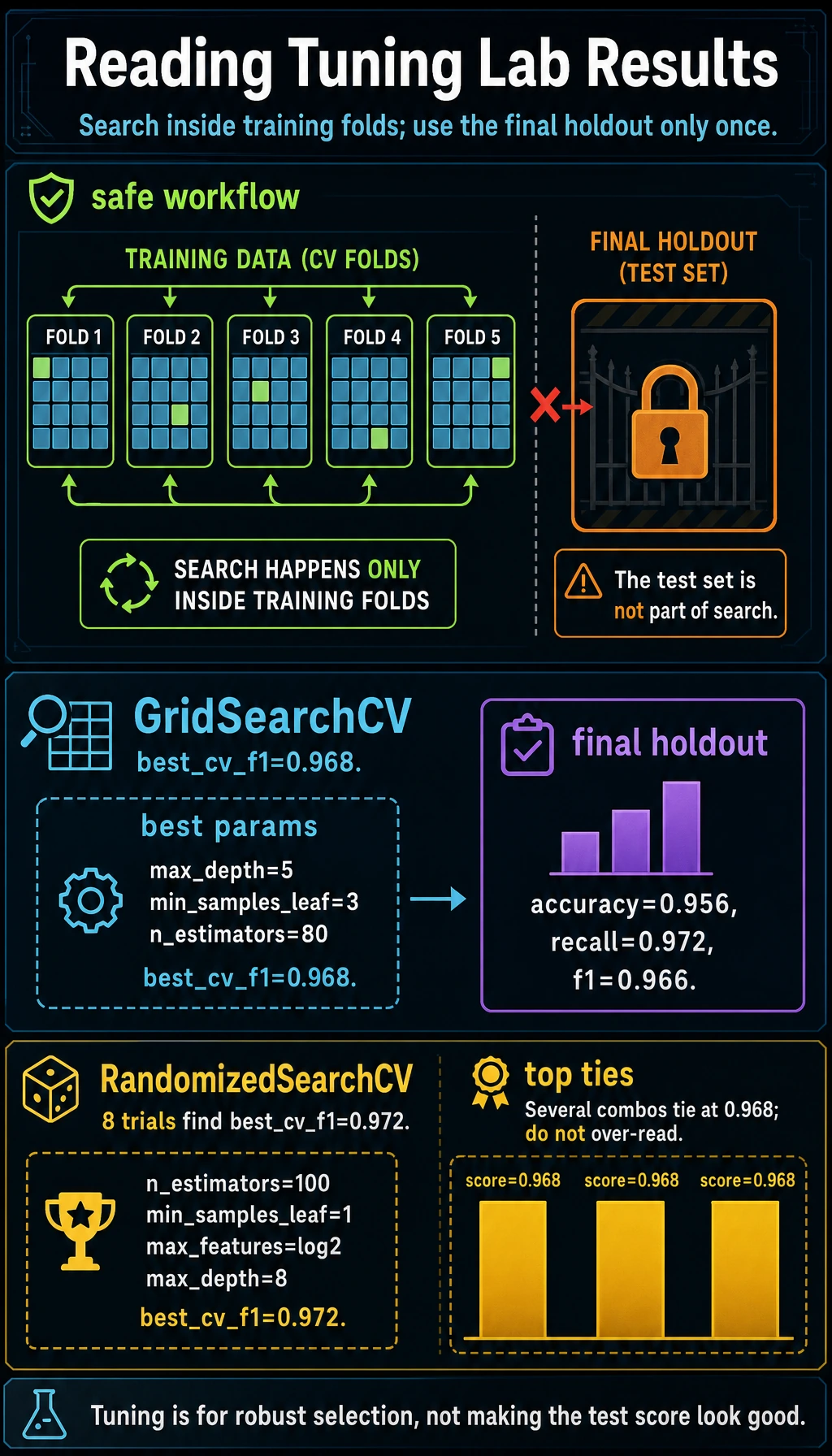

Parameters vs Hyperparameters
Random Forest learns split rules from data. Those learned split rules are parameters.
You choose settings such as:
n_estimators;max_depth;min_samples_leaf;max_features.
Those are hyperparameters. They shape how learning happens.
Grid Search
Grid search tries every combination you list:
param_grid={
"n_estimators": [80, 160],
"max_depth": [3, 5, None],
"min_samples_leaf": [1, 3],
}
This grid has 2 x 3 x 2 = 12 combinations. With 5-fold CV, that means 60 model fits.
Use grid search when:
- the search space is small;
- you understand which values are plausible;
- you want a simple, repeatable baseline.
Random Search
Random search samples a limited number of combinations from a larger space:
n_iter=8
In the lab, it tried only 8 combinations but searched a wider space and found a slightly higher CV F1:
best_cv_f1=0.972
Use random search when:
- there are many hyperparameters;
- training is expensive;
- you want to explore before designing a narrower grid.
Final Holdout
The final holdout is the part we did not use in CV search:
X_train, X_final, y_train, y_final = train_test_split(...)
After the search chooses the best settings, we evaluate once:
final accuracy=0.956 recall=0.972 f1=0.966
Do not keep changing the grid after looking at the final holdout. If you do, it stops being a final holdout and becomes part of tuning.
Reading Search Results
The top grid results are very close:
score=0.968 params={'max_depth': 5, 'min_samples_leaf': 3, 'n_estimators': 80}
score=0.968 params={'max_depth': 5, 'min_samples_leaf': 3, 'n_estimators': 160}
When scores tie, prefer the simpler or cheaper model. More trees or deeper trees are not automatically better.
Practical Tuning Strategy
| Stage | Action |
|---|---|
| Start | Build a simple baseline with default settings |
| Diagnose | Check bias/variance and metric choice |
| First search | Use a small grid around important parameters |
| Wider search | Use random search when combinations explode |
| Final check | Evaluate once on untouched holdout |
| Production | monitor drift and retrain policy |
For experienced readers: Bayesian tools such as Optuna are useful when each trial is expensive or the search space is large. They are not a replacement for clean validation design.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| Search takes too long | grid too large | reduce candidates, use random search |
| CV score rises but final holdout drops | over-tuning | simplify search, keep a fresh holdout |
| Best model is much more complex | metric difference is tiny | choose cheaper/simpler model |
| Different runs choose different params | unstable data or small folds | use repeated CV or inspect variance |
| Tuning does not help | model class or features are limiting | improve features or model family first |
Practice
- Change scoring from
"f1"to"recall". Which parameters change? - Add
max_depth=10to the grid. Does it improve CV score? - Increase
n_iterfrom8to16. Does random search improve enough to justify the cost? - Print
mean_fit_timefromcv_results_and choose a cheaper model when scores tie. - Add a final untouched test set to one earlier lesson that currently uses only CV.
Pass Check
You are done when you can explain:
- hyperparameters are chosen before training;
- grid search is exhaustive over a small listed space;
- random search is useful when the space is larger;
- final holdout must not be used for repeated tuning;
- tuning cannot rescue bad features or the wrong validation design.