5.2.6 SVM: Maximum Margin and Kernel Methods
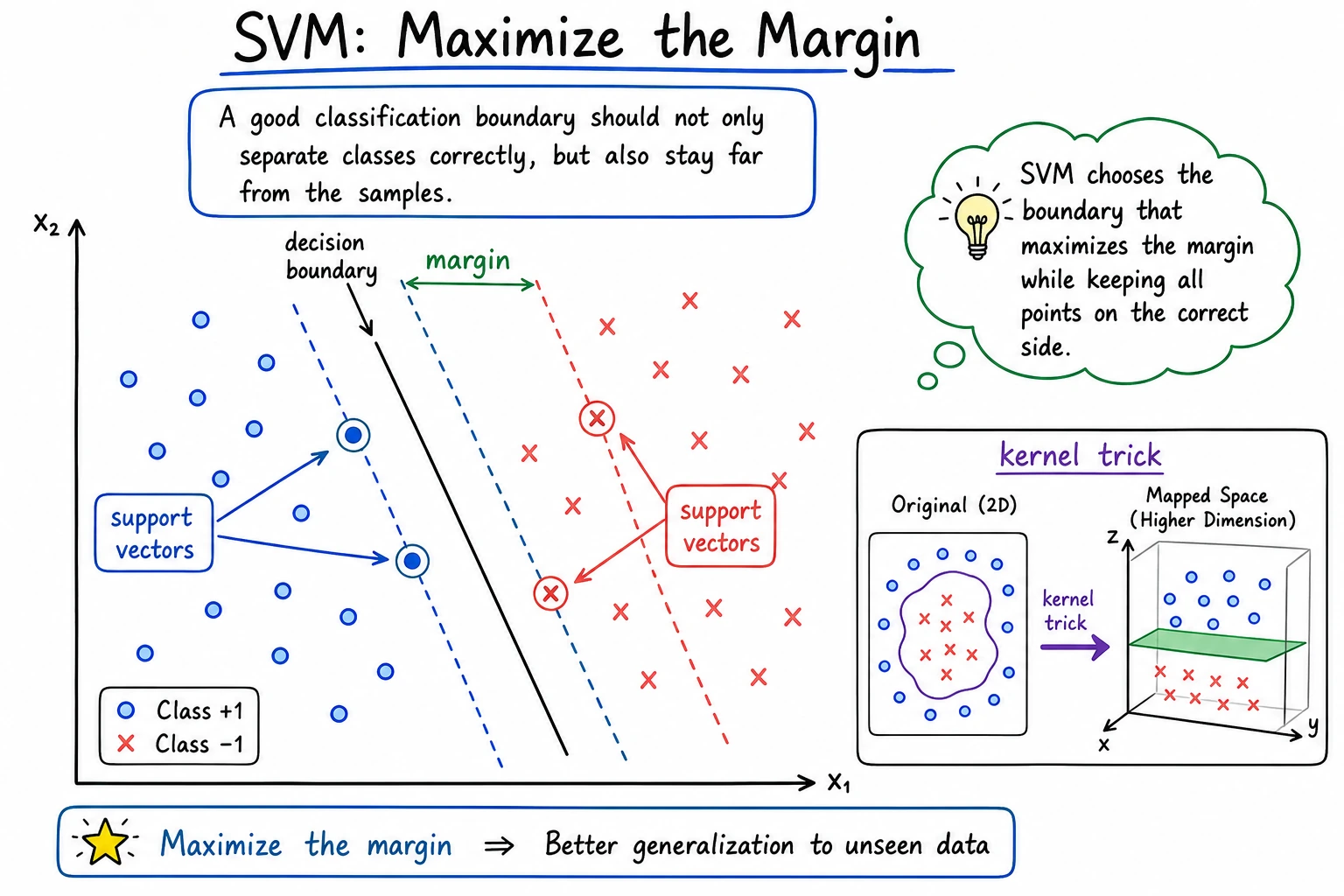
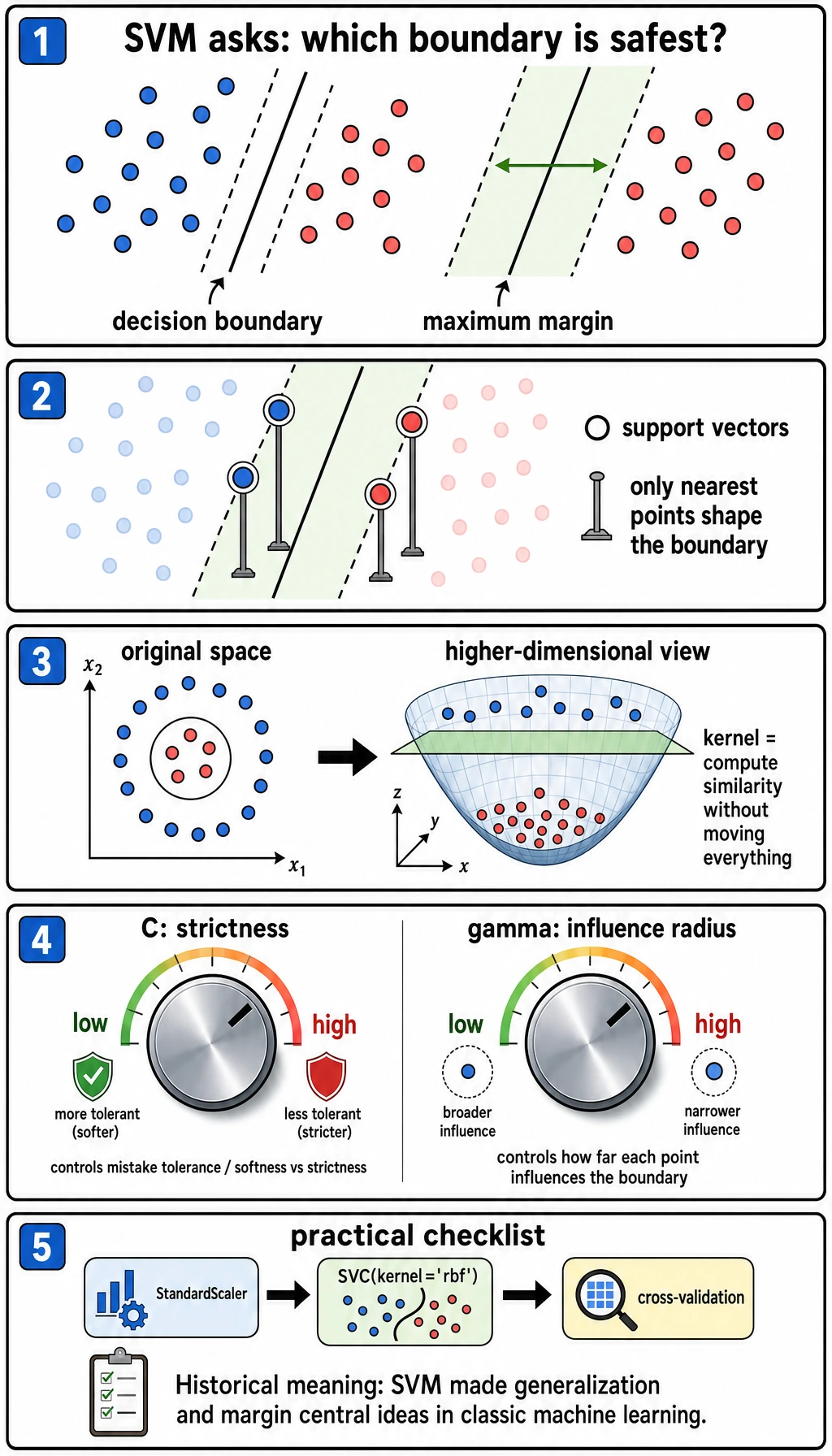
SVM is not always the first production model today, but it is still one of the clearest ways to learn margin, kernel, and distance-sensitive modeling.
What You Will Build
This lesson turns SVM into a small lab. You will:
- compare
linearandrbfkernels on a curved dataset; - prove why
StandardScalermatters for SVM; - tune
Candgammaand inspect support vector counts; - learn when SVM is worth trying and when ensembles are usually easier.
The practical sentence to remember:
SVM does not only ask "did I classify this correctly?" It asks "can I place the boundary with enough room around the closest samples?"
Keyword Decoder
| Term | Practical meaning |
|---|---|
SVM | Support Vector Machine, a classifier that searches for a large-margin boundary |
margin | Distance between the boundary and the closest samples |
support vector | A training sample close enough to shape the boundary |
kernel | A similarity function that lets SVM create nonlinear boundaries |
RBF | Radial Basis Function, a common nonlinear kernel |
C | Mistake penalty; larger C tries harder to fit training points |
gamma | Local influence radius for the RBF kernel; larger values create more local boundaries |
SVC | sklearn's Support Vector Classifier |
Setup
python -m pip install -U scikit-learn
SVM is sensitive to feature scale, so the examples use Pipeline(StandardScaler(), SVC(...)). This is not decoration; it is part of the model workflow.
Run the Complete Lab
Create svm_lab.py:
from itertools import product
from sklearn.datasets import make_moons
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVC
X, y = make_moons(n_samples=400, noise=0.25, random_state=42)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42, stratify=y
)
print("kernel_comparison")
for kernel in ["linear", "rbf"]:
model = make_pipeline(StandardScaler(), SVC(kernel=kernel, C=1.0, gamma="scale"))
model.fit(X_train, y_train)
svc = model.named_steps["svc"]
print(
f"kernel={kernel:<6} "
f"accuracy={accuracy_score(y_test, model.predict(X_test)):.3f} "
f"support_vectors={int(svc.n_support_.sum())}"
)
print("scaling_check")
X_bad_scale = X.copy()
X_bad_scale[:, 1] *= 100
X_train2, X_test2, y_train2, y_test2 = train_test_split(
X_bad_scale, y, test_size=0.25, random_state=42, stratify=y
)
raw = SVC(kernel="rbf", C=1.0, gamma="scale")
raw.fit(X_train2, y_train2)
scaled = make_pipeline(StandardScaler(), SVC(kernel="rbf", C=1.0, gamma="scale"))
scaled.fit(X_train2, y_train2)
print(f"without_scaling={accuracy_score(y_test2, raw.predict(X_test2)):.3f}")
print(f"with_scaling={accuracy_score(y_test2, scaled.predict(X_test2)):.3f}")
print("c_gamma_lab")
for C, gamma in product([0.1, 1.0, 10.0], [0.1, 1.0]):
model = make_pipeline(StandardScaler(), SVC(kernel="rbf", C=C, gamma=gamma))
model.fit(X_train, y_train)
svc = model.named_steps["svc"]
print(
f"C={C:<4} gamma={gamma:<3} "
f"accuracy={accuracy_score(y_test, model.predict(X_test)):.3f} "
f"support_vectors={int(svc.n_support_.sum())}"
)
Run it:
python svm_lab.py
Expected output:
kernel_comparison
kernel=linear accuracy=0.920 support_vectors=125
kernel=rbf accuracy=0.950 support_vectors=98
scaling_check
without_scaling=0.880
with_scaling=0.950
c_gamma_lab
C=0.1 gamma=0.1 accuracy=0.940 support_vectors=187
C=0.1 gamma=1.0 accuracy=0.960 support_vectors=173
C=1.0 gamma=0.1 accuracy=0.950 support_vectors=134
C=1.0 gamma=1.0 accuracy=0.930 support_vectors=87
C=10.0 gamma=0.1 accuracy=0.960 support_vectors=111
C=10.0 gamma=1.0 accuracy=0.920 support_vectors=57
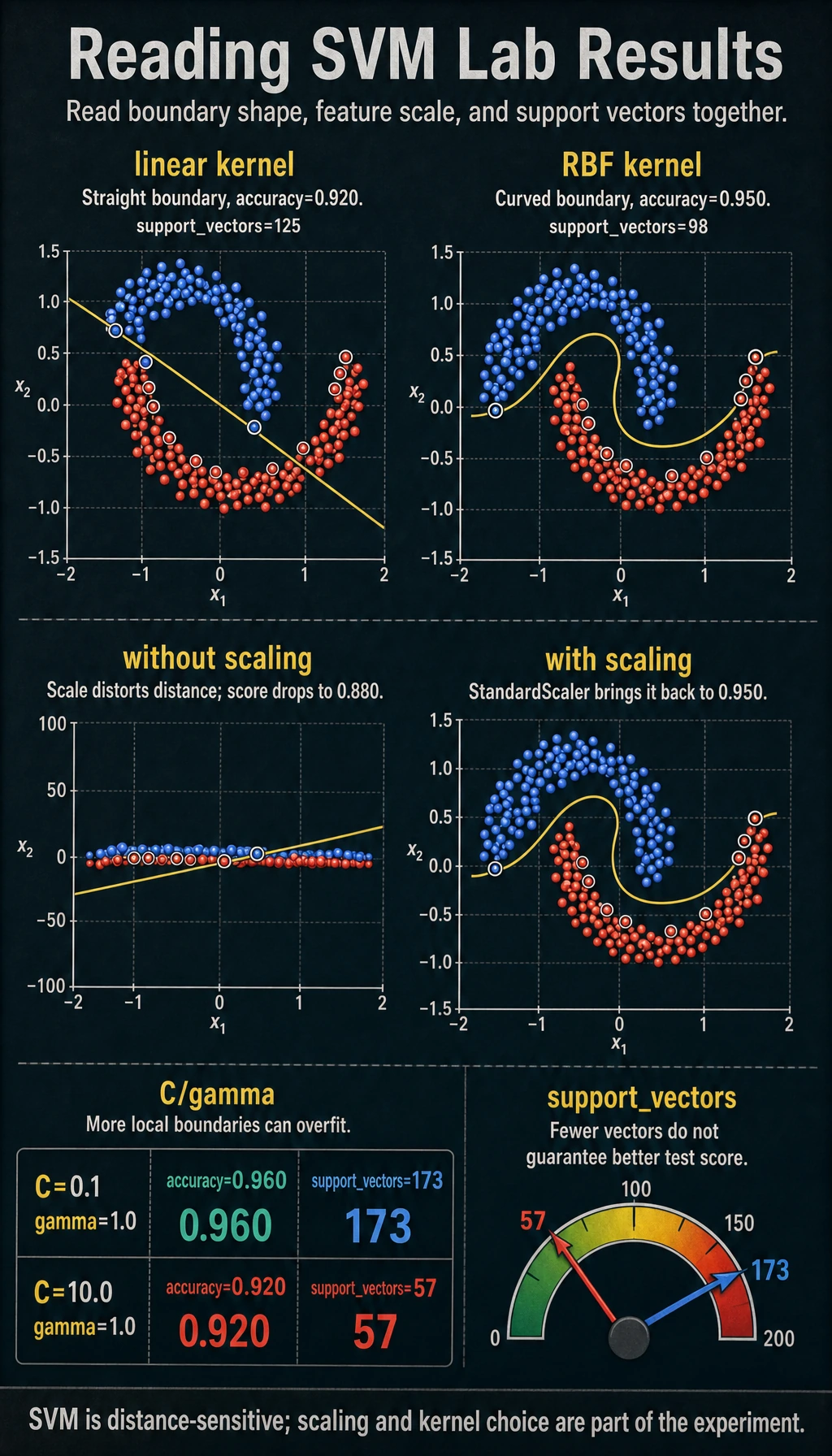
Read the Kernel Result
The curved make_moons dataset is intentionally hard for a straight boundary:
kernel=linear accuracy=0.920 support_vectors=125
kernel=rbf accuracy=0.950 support_vectors=98
The linear kernel asks for a straight separating line. The rbf kernel compares local similarity, so it can create a curved boundary. Use this simple rule:
| Situation | First SVM choice |
|---|---|
| Boundary looks roughly straight | kernel="linear" |
| Boundary is curved and the dataset is not huge | kernel="rbf" |
| You have many rows or many features | Try logistic regression, linear SVM, or tree ensembles first |
Why Scaling Is Not Optional
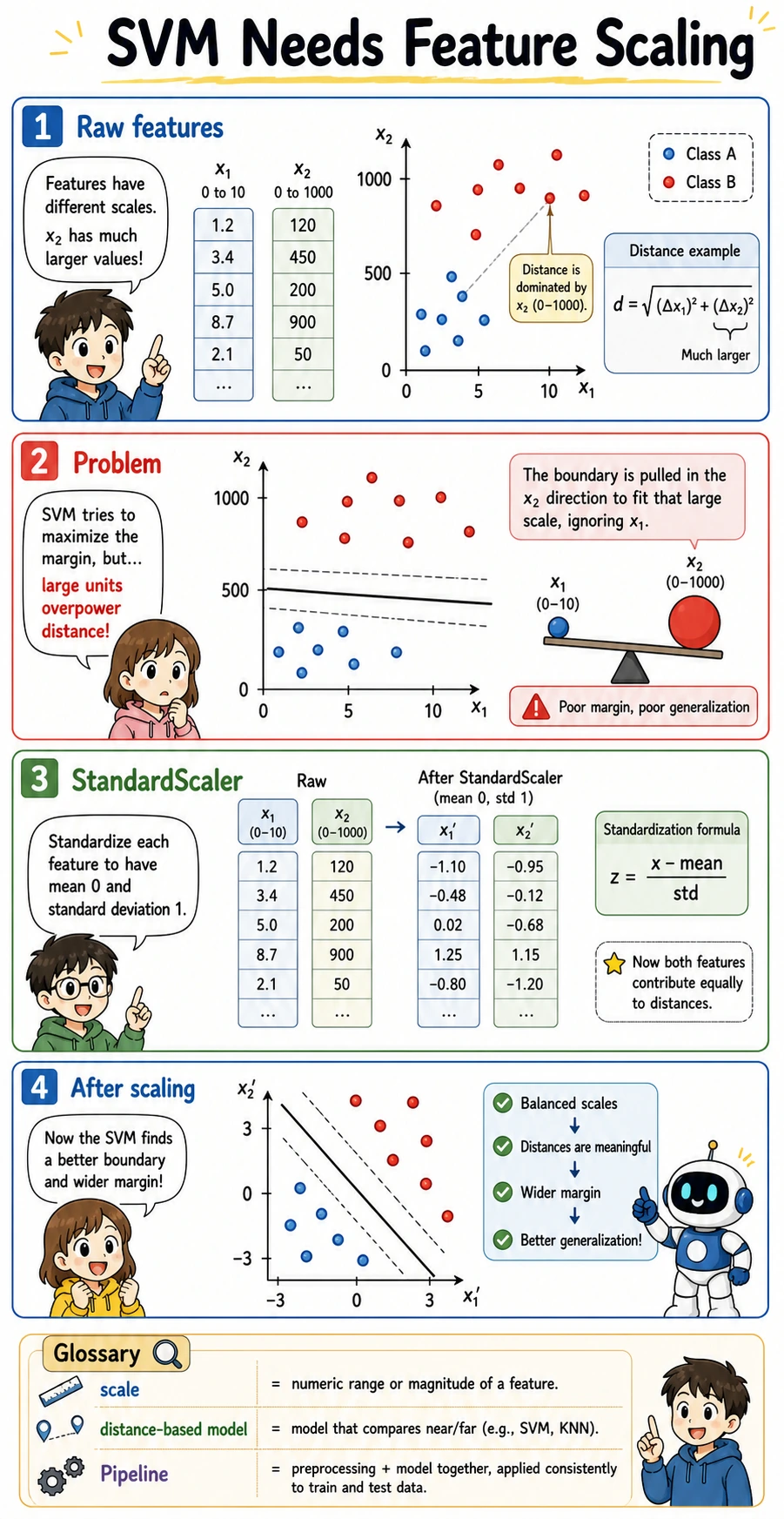
SVM relies on distances and similarities. If one feature has values around 0-1 and another has values around 0-1000, the larger-scale feature can dominate the boundary even when it is not more meaningful.
The lab makes that problem visible:
without_scaling=0.880
with_scaling=0.950
This is why StandardScaler should live inside a Pipeline: the scaler is fitted only on the training fold, then applied safely to validation/test data.
Understand C and gamma
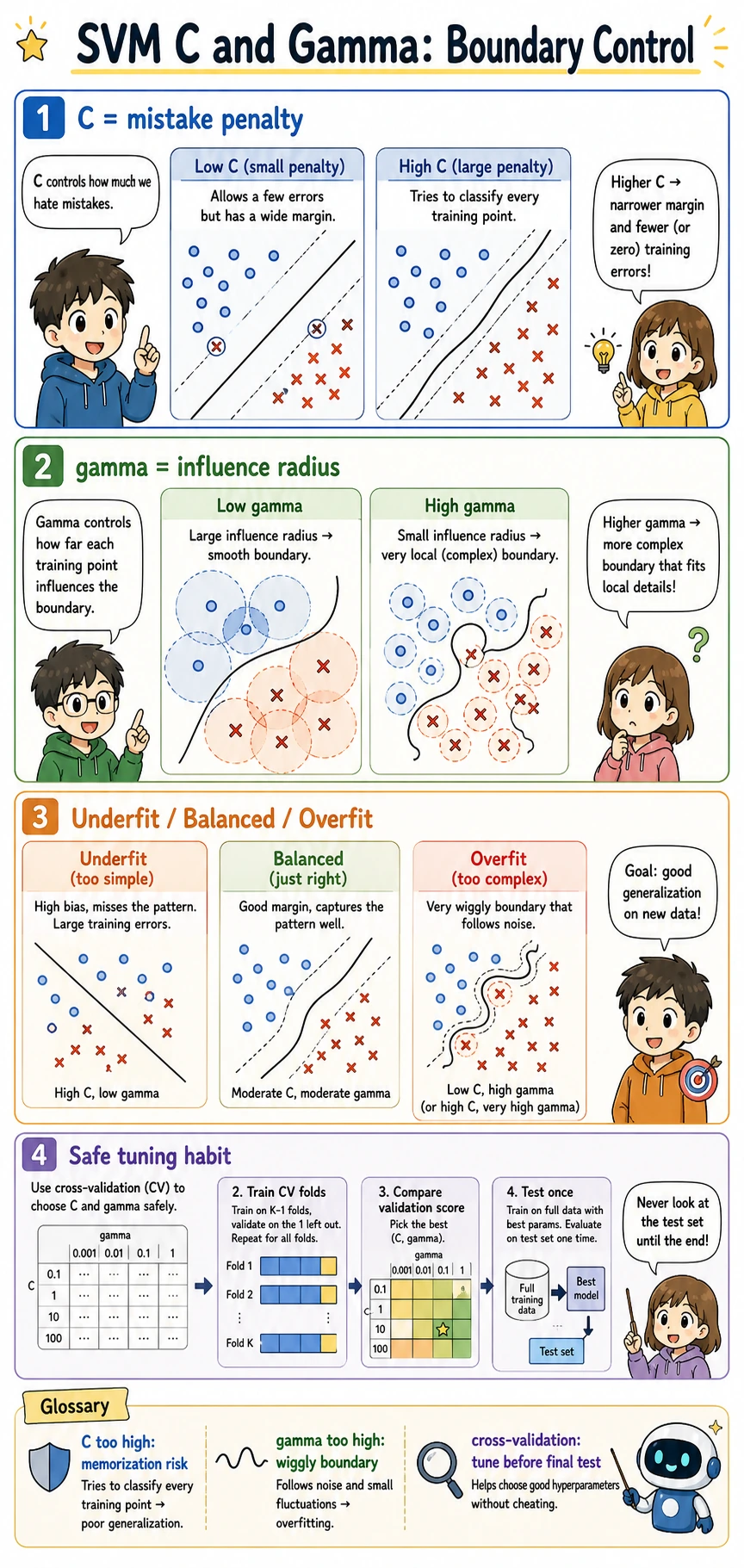

C and gamma control different parts of the boundary:
| Parameter | If too small | If too large |
|---|---|---|
C | allows more mistakes; wider, smoother margin | chases training points more aggressively |
gamma | influence is broad; boundary may be too smooth | influence is local; boundary can become wiggly |
Read the output with two signals:
C=0.1 gamma=1.0 accuracy=0.960 support_vectors=173
C=10.0 gamma=1.0 accuracy=0.920 support_vectors=57
The second model uses fewer support vectors, but its test accuracy is worse. Fewer support vectors is not automatically better. It can mean the model is using a sharper boundary that generalizes poorly.
For experienced readers: tune C and gamma with cross-validation, and compare against logistic regression and ensemble baselines. Do not select SVM from one train-test split.
Support Vectors in Practice
Support vectors are the points close enough to the boundary to matter. They are useful for intuition:
- many support vectors can mean the boundary is uncertain or the margin is soft;
- very few support vectors with poor test score can signal an overly sharp boundary;
- support vector count is a diagnostic hint, not a final metric.
If you need calibrated probabilities, remember that SVC(probability=True) adds an extra calibration step and costs more training time. Often it is cleaner to use CalibratedClassifierCV when probability quality matters.
When to Use SVM
SVM is worth trying when:
- the dataset is small to medium sized;
- features are numeric and well-scaled;
- you need a strong nonlinear classifier without building a neural network;
- you want to understand margin-based classification.
Prefer other models when:
- you need fast training on very large data;
- you have many categorical features that need heavy preprocessing;
- probability calibration is central to the product;
- tree ensembles already perform better with less tuning.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| SVM performs much worse than expected | features are not scaled | use StandardScaler inside Pipeline |
| Training is slow | RBF SVM does not scale well to large datasets | try linear models, LinearSVC, or ensembles |
| Boundary seems too wiggly | gamma or C is too large | lower gamma, lower C, use cross-validation |
| Model misses curved patterns | using linear when boundary is nonlinear | compare with kernel="rbf" |
| Need reliable probabilities | raw SVM scores are not calibrated probabilities | use calibration and check probability metrics |
Practice
- Change
noiseinmake_moons()from0.25to0.1and0.4. Which settings make SVM easier or harder? - Add
gamma=5.0to the grid. What happens to accuracy and support vector count? - Replace
SVCwithLinearSVCfor the linear case. What changes in available attributes? - Run logistic regression on the same dataset and compare it with RBF SVM.
- Use cross-validation to pick
Candgammainstead of trusting one split.
Pass Check
You are done when you can explain:
- SVM searches for a boundary with a large margin;
- support vectors are the boundary-critical training points;
- RBF kernel can model curved boundaries;
- scaling is essential because SVM uses distances;
Candgammamust be tuned together, preferably with cross-validation.