5.2.4 Decision Trees
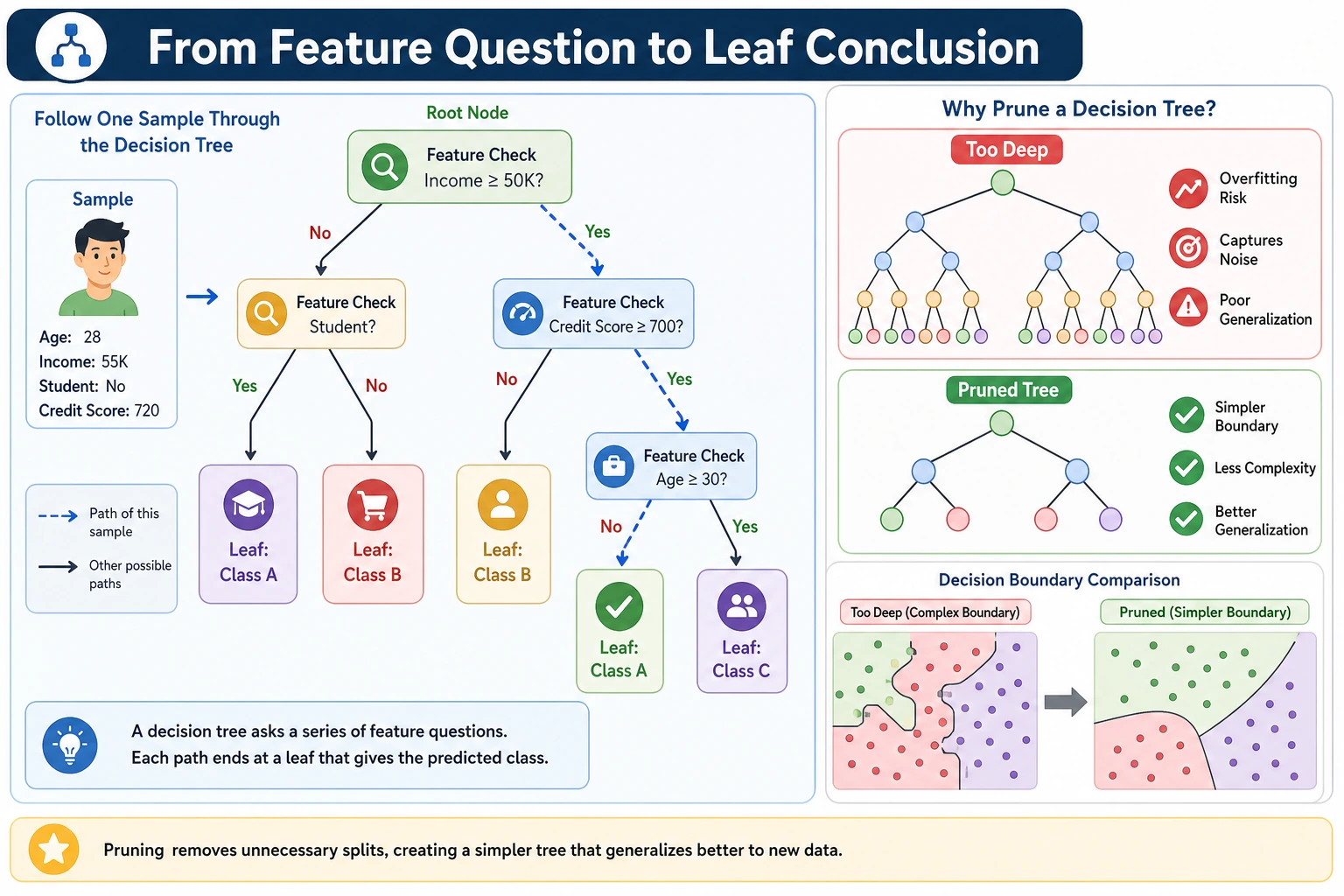
A decision tree is a model made of questions. It is easy to read because each prediction follows a path of rules, but it can also overfit quickly when the rules become too detailed.
What You Will Build
In this lesson you will run one script that shows:
- how tree depth changes train/test accuracy;
- how to print readable tree rules;
- how feature importance is computed from splits;
- how post-pruning with
ccp_alphachanges the number of leaves; - how a regression tree makes step-like numeric predictions.
Read the maps first. A tree is not just "if-else"; it is "if-else plus a scoring rule plus complexity control."
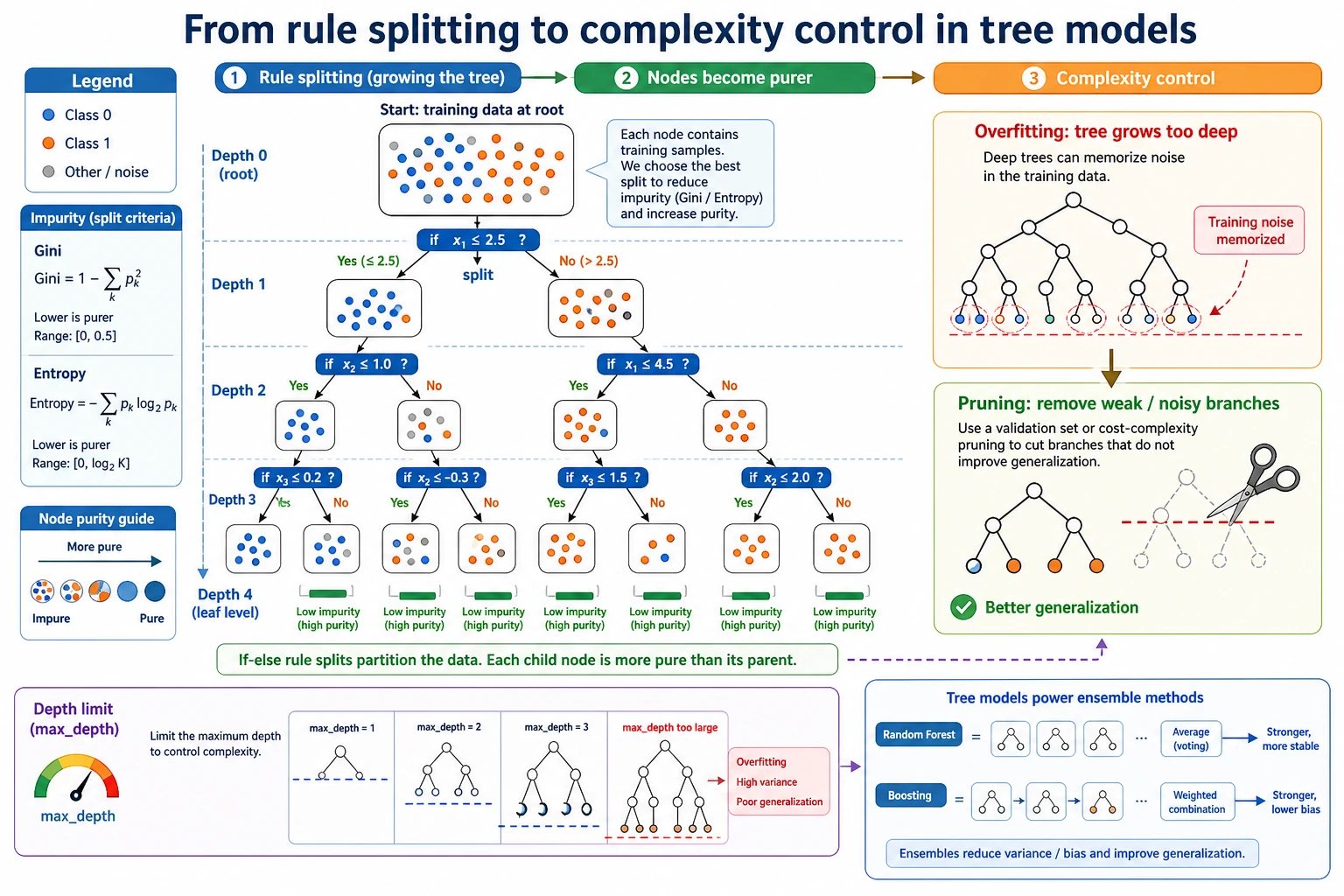
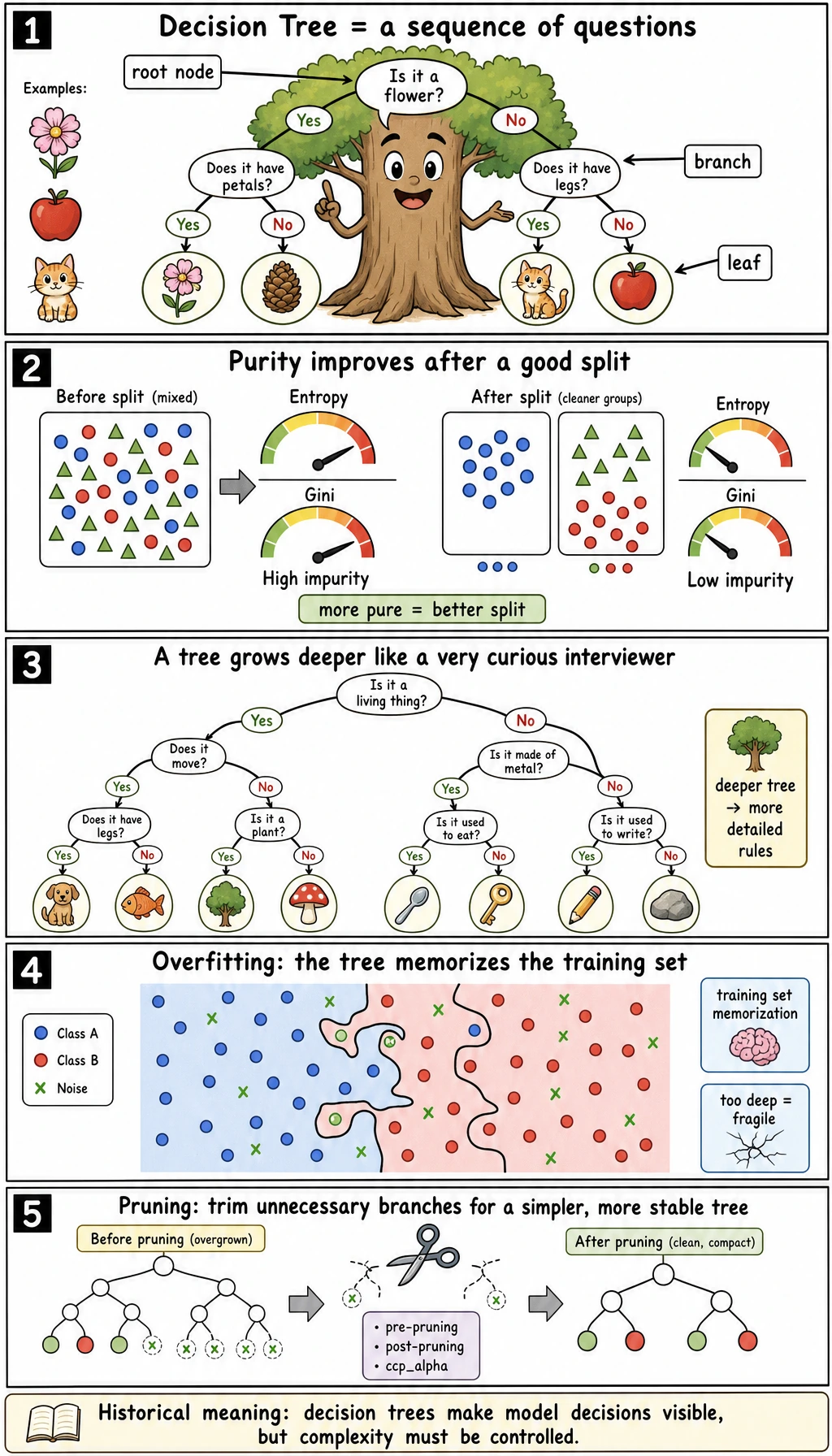
Setup
python -m pip install -U scikit-learn
This lesson uses sklearn's CART-style DecisionTreeClassifier and DecisionTreeRegressor. CART means Classification and Regression Trees: the same tree idea can handle both labels and numbers.
Run the Complete Lab
Create decision_tree_lab.py:
from sklearn.datasets import load_diabetes, load_iris
from sklearn.metrics import accuracy_score, mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor, export_text
iris = load_iris()
X = iris.data[:, 2:4] # petal length and petal width, easier to read
y = iris.target
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42, stratify=y
)
print("classification_depth_lab")
for depth in [1, 2, 3, None]:
tree = DecisionTreeClassifier(max_depth=depth, min_samples_leaf=3, random_state=42)
tree.fit(X_train, y_train)
train_acc = accuracy_score(y_train, tree.predict(X_train))
test_acc = accuracy_score(y_test, tree.predict(X_test))
print(
f"max_depth={str(depth):<4} "
f"train={train_acc:.3f} test={test_acc:.3f} "
f"leaves={tree.get_n_leaves()} depth={tree.get_depth()}"
)
best_tree = DecisionTreeClassifier(max_depth=3, min_samples_leaf=3, random_state=42)
best_tree.fit(X_train, y_train)
print("tree_rules")
print(export_text(best_tree, feature_names=["petal length", "petal width"], decimals=2, max_depth=3))
print("feature_importance")
for name, value in zip(["petal length", "petal width"], best_tree.feature_importances_):
print(f"- {name}: {value:.3f}")
print("pruning_lab")
path = DecisionTreeClassifier(random_state=42).cost_complexity_pruning_path(X_train, y_train)
for alpha in path.ccp_alphas[[0, 1, -2]]:
pruned = DecisionTreeClassifier(random_state=42, ccp_alpha=float(alpha))
pruned.fit(X_train, y_train)
print(
f"ccp_alpha={alpha:.4f} "
f"test={accuracy_score(y_test, pruned.predict(X_test)):.3f} "
f"leaves={pruned.get_n_leaves()}"
)
print("regression_tree_lab")
diabetes = load_diabetes()
X_train, X_test, y_train, y_test = train_test_split(
diabetes.data, diabetes.target, test_size=0.25, random_state=42
)
for depth in [2, 4, None]:
reg = DecisionTreeRegressor(max_depth=depth, min_samples_leaf=10, random_state=42)
reg.fit(X_train, y_train)
pred = reg.predict(X_test)
print(f"max_depth={str(depth):<4} mae={mean_absolute_error(y_test, pred):.1f} leaves={reg.get_n_leaves()}")
Run it:
python decision_tree_lab.py
Expected output:
classification_depth_lab
max_depth=1 train=0.670 test=0.658 leaves=2 depth=1
max_depth=2 train=0.964 test=0.947 leaves=3 depth=2
max_depth=3 train=0.982 test=0.974 leaves=5 depth=3
max_depth=None train=0.982 test=0.974 leaves=5 depth=3
tree_rules
|--- petal length <= 2.45
| |--- class: 0
|--- petal length > 2.45
| |--- petal width <= 1.70
| | |--- petal length <= 4.95
| | | |--- class: 1
| | |--- petal length > 4.95
| | | |--- class: 2
| |--- petal width > 1.70
| | |--- petal length <= 4.95
| | | |--- class: 2
| | |--- petal length > 4.95
| | | |--- class: 2
feature_importance
- petal length: 0.588
- petal width: 0.412
pruning_lab
ccp_alpha=0.0000 test=0.921 leaves=7
ccp_alpha=0.0067 test=0.921 leaves=5
ccp_alpha=0.2636 test=0.658 leaves=2
regression_tree_lab
max_depth=2 mae=47.3 leaves=4
max_depth=4 mae=44.4 leaves=14
max_depth=None mae=48.7 leaves=25
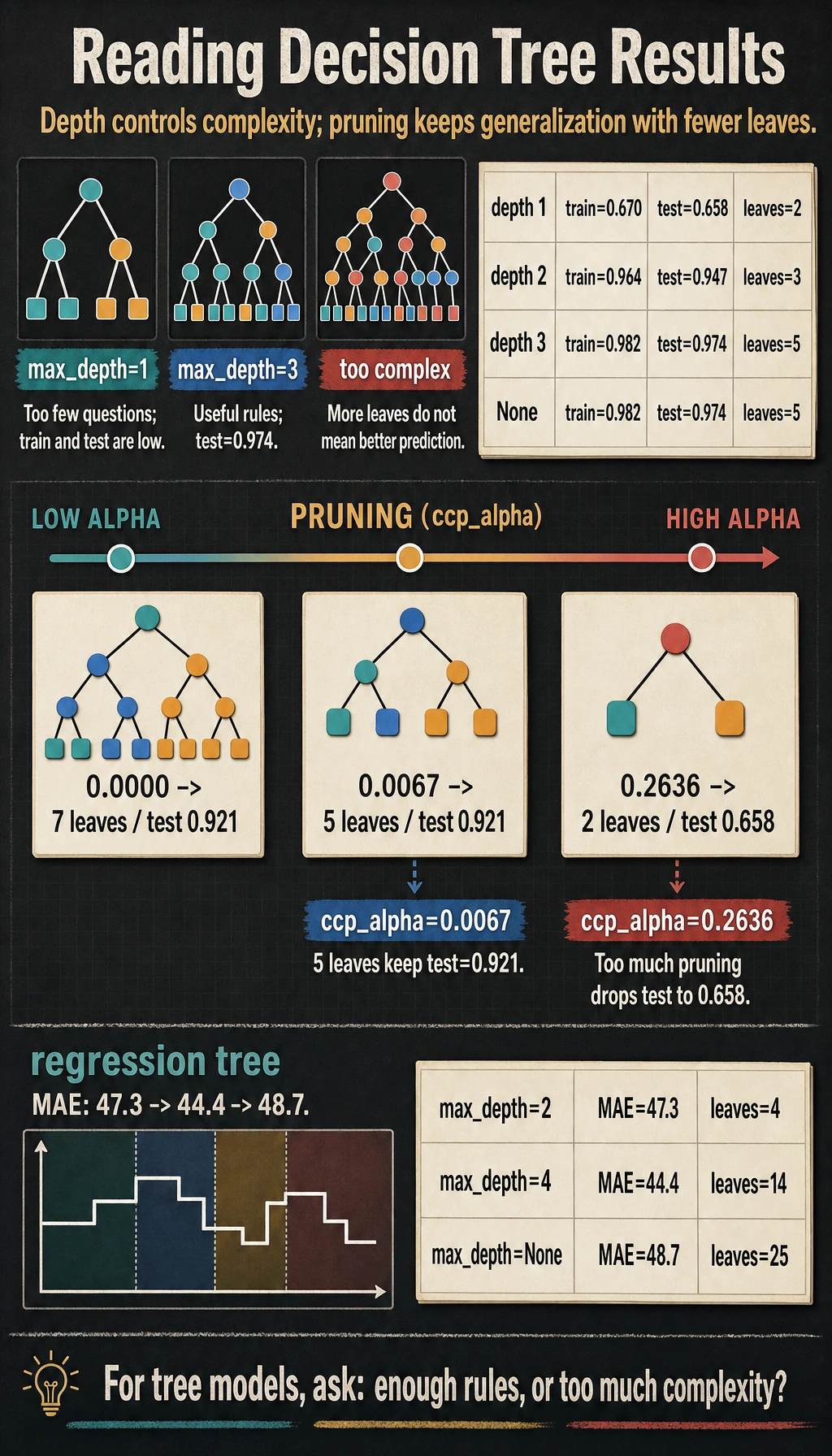
Read the Output
The first block is the most important part:
max_depth=1 train=0.670 test=0.658 leaves=2 depth=1
max_depth=3 train=0.982 test=0.974 leaves=5 depth=3
max_depth=1 asks only one question, so the model is too simple. max_depth=3 asks a few follow-up questions and performs much better. On this tiny dataset, max_depth=None does not grow deeper because min_samples_leaf=3 and the data are already simple enough.
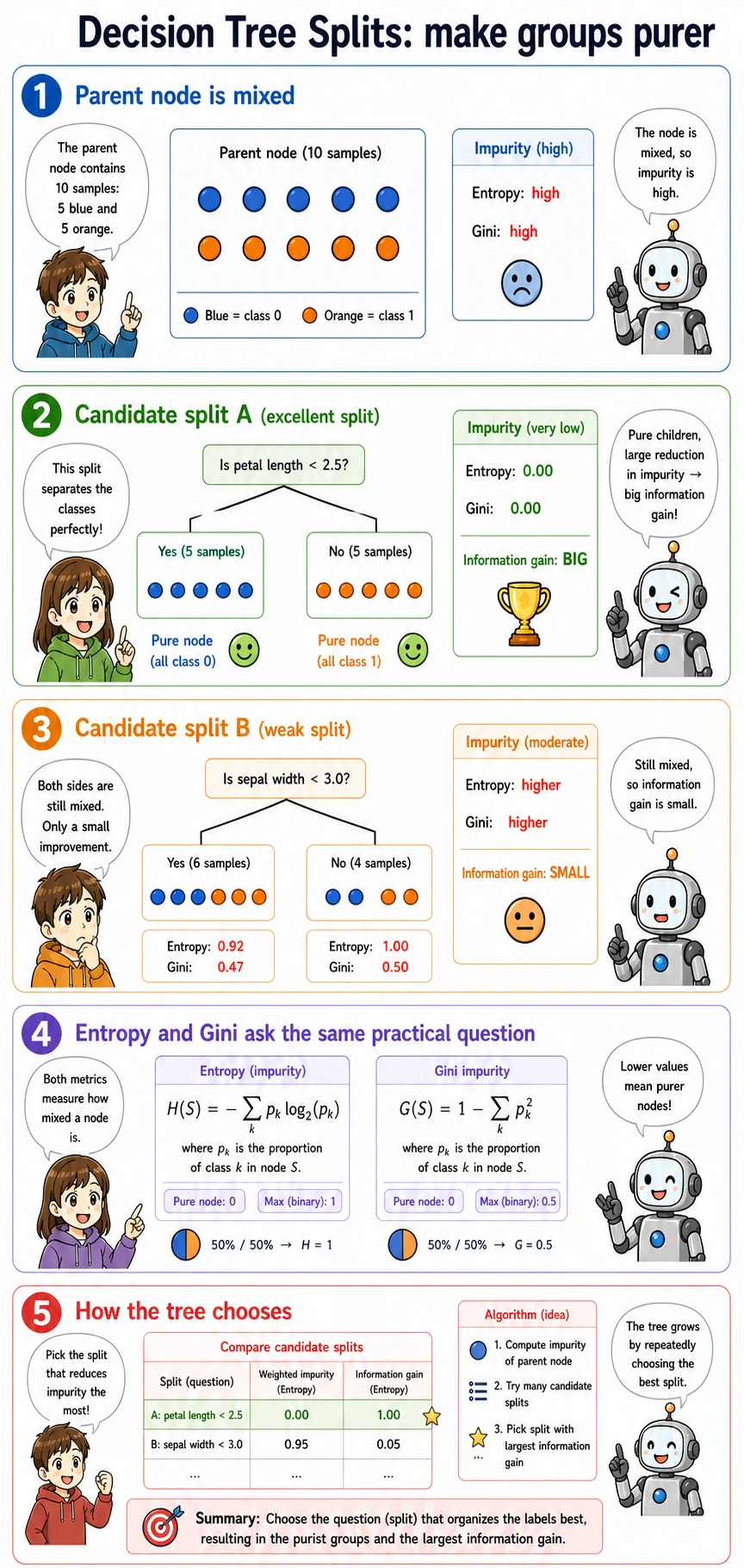

At each node, the tree searches for a question like:
petal length <= 2.45?
The chosen split should make the child nodes cleaner than the parent node. Cleaner means the labels inside a node are less mixed.
Gini, Entropy, and Information Gain
You do not need to memorize every formula on first pass. Remember the job:
| Term | Practical meaning |
|---|---|
Gini | How mixed the labels are in a node; sklearn's default for classification trees |
entropy | Another mixedness score, connected to information theory |
information gain | How much mixedness drops after a split |
criterion | The setting that chooses the scoring rule, such as criterion="gini" or criterion="entropy" |
Use gini first unless you have a reason to compare. In many tabular projects, tuning depth, leaf size, and pruning matters more than switching from Gini to entropy.
Complexity Control
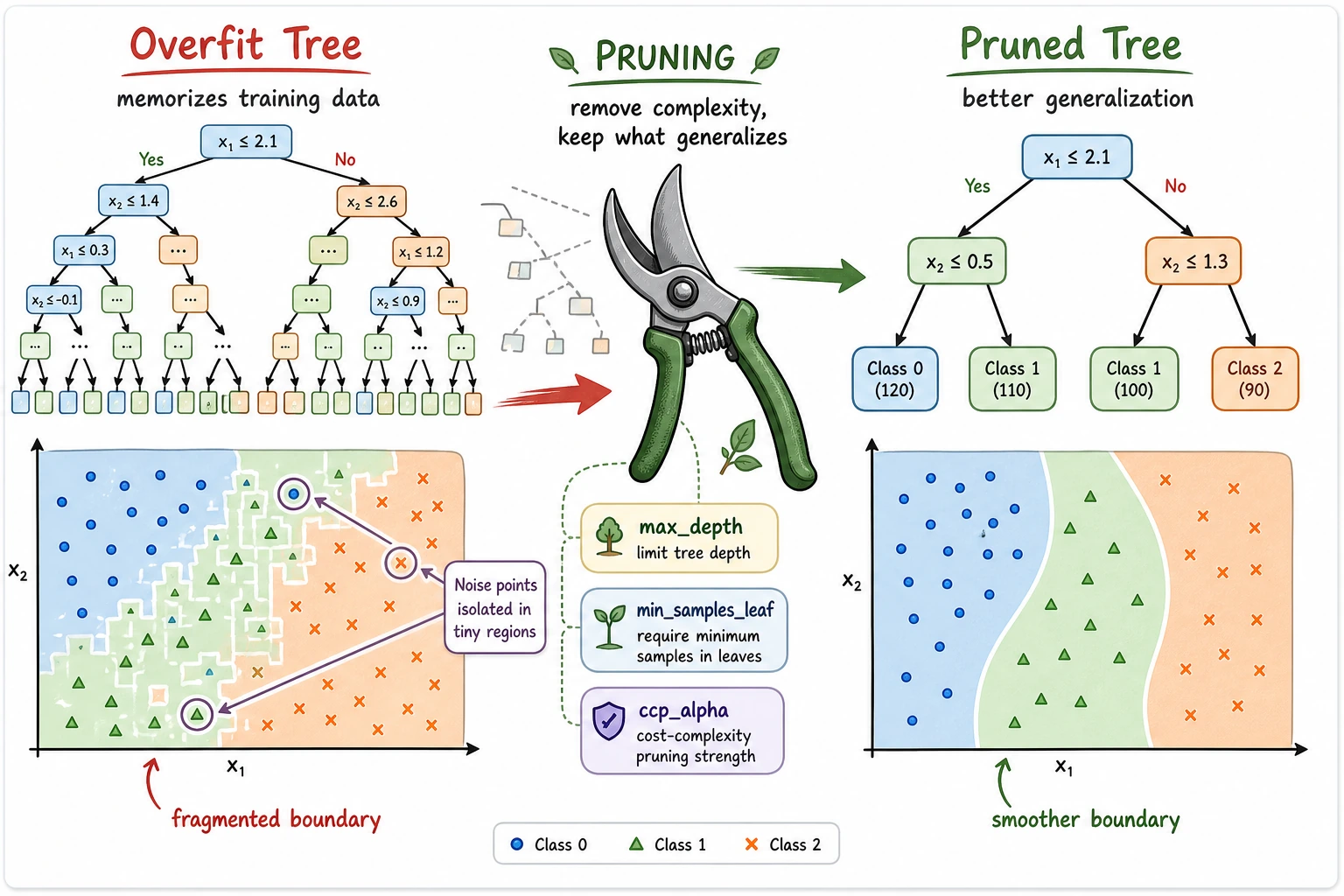
The practical tuning order is:
- Set
max_depthto stop the tree from growing too deep. - Set
min_samples_leafso each leaf has enough examples. - Use
ccp_alphafor post-pruning after a full tree is grown.
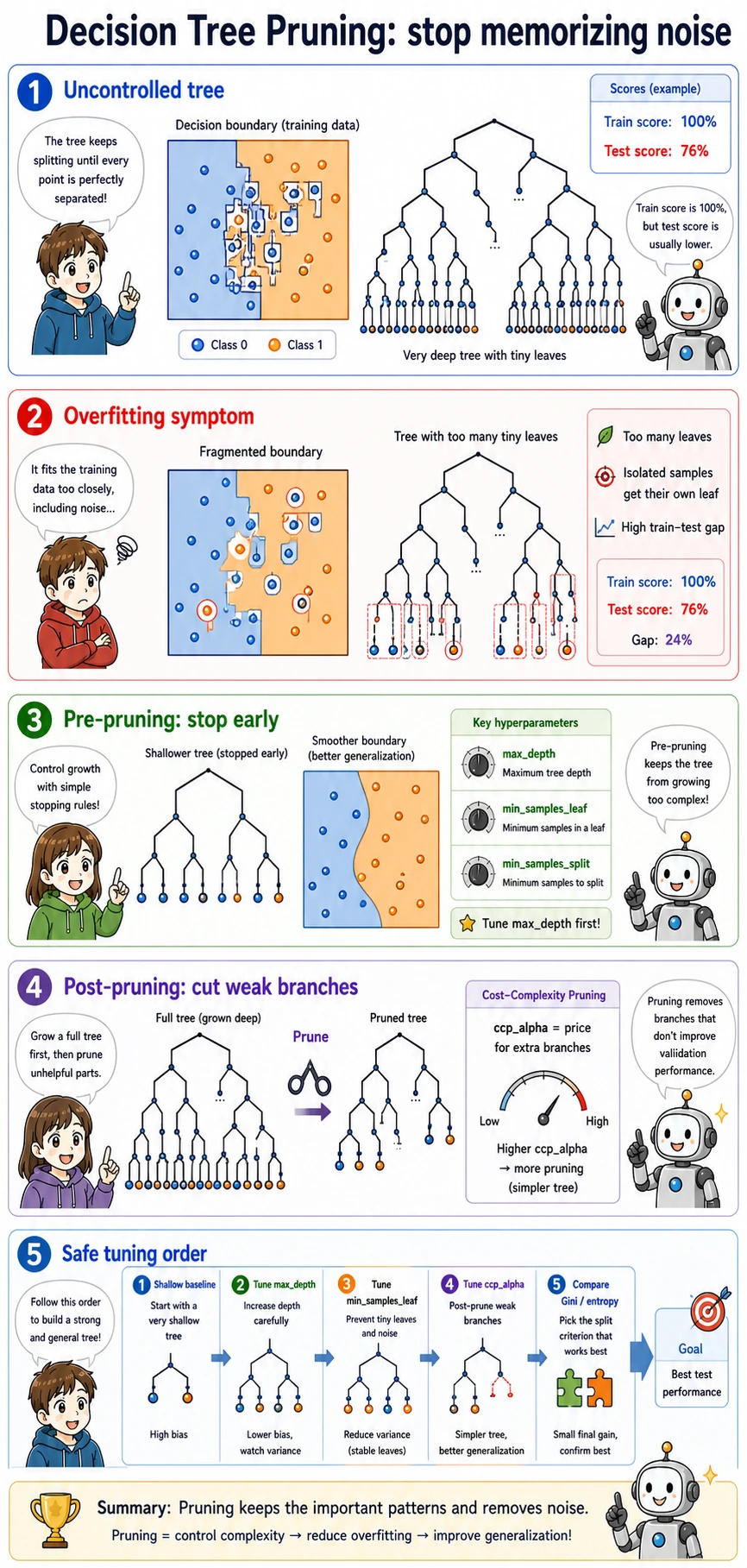
The pruning output shows the trade-off:
ccp_alpha=0.0000 test=0.921 leaves=7
ccp_alpha=0.0067 test=0.921 leaves=5
ccp_alpha=0.2636 test=0.658 leaves=2
A little pruning kept the same test score with fewer leaves. Too much pruning collapsed the tree into only two leaves and lost useful rules.
Interpretability
export_text() prints the path a sample will follow. This is useful when you need to explain a prediction to a teammate:
|--- petal length <= 2.45
| |--- class: 0
Feature importance is also useful, but read it carefully:
- it tells you which features reduced impurity most in this fitted tree;
- it can favor features with many possible split points;
- correlated features can share or hide importance;
- it is not the same as causal importance.
For stronger interpretation, compare tree importance with permutation importance later.
Regression Trees
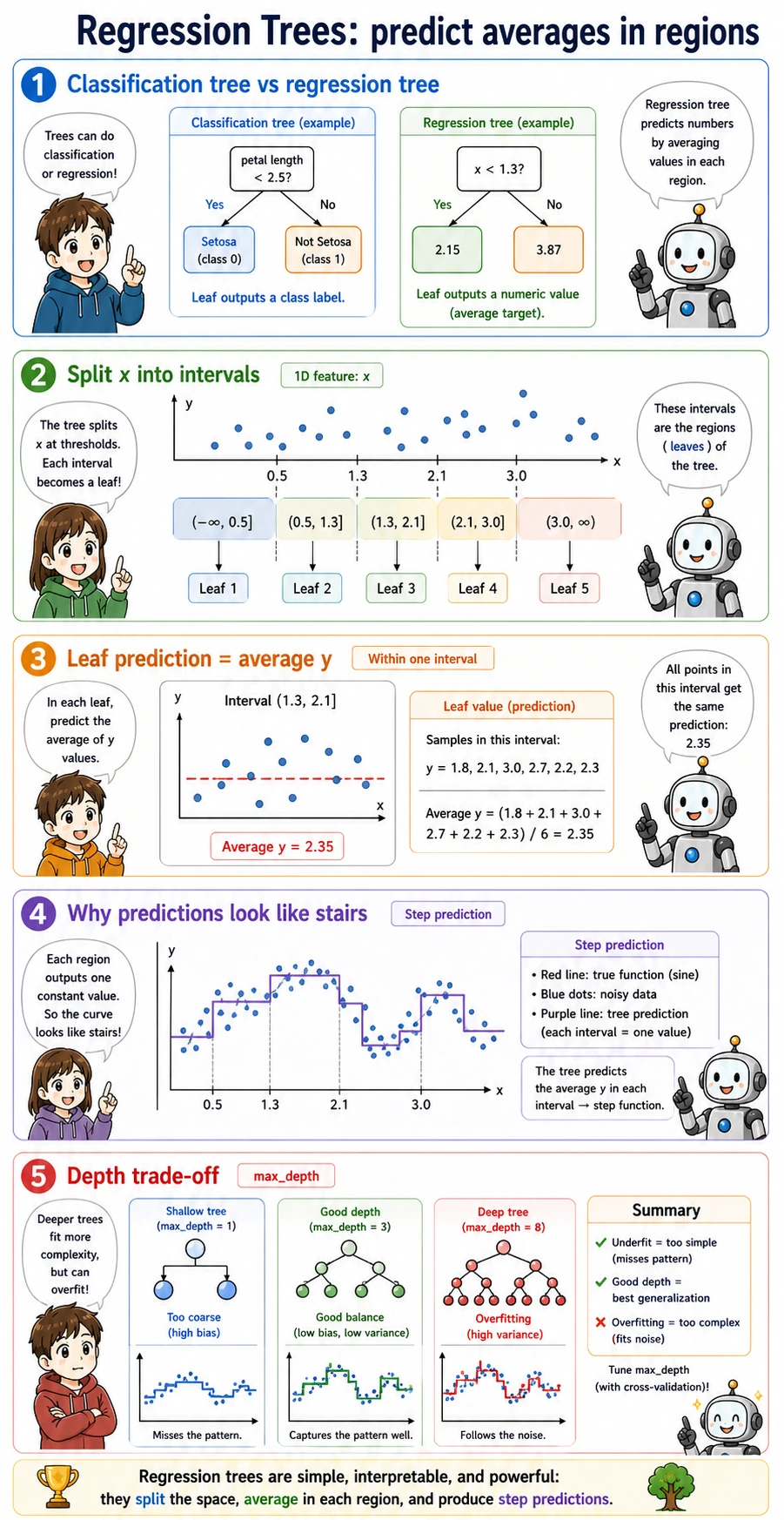

A regression tree predicts numbers, but the idea is the same: split the feature space into regions, then output the average target value in each leaf.
That is why regression tree predictions often look like steps rather than a smooth line. In the lab:
max_depth=4 mae=44.4 leaves=14
max_depth=None mae=48.7 leaves=25
The deeper tree has more leaves, but its test MAE is worse. More rules do not automatically mean better generalization.
When to Use a Single Decision Tree
Use a single tree when you need:
- a quick, explainable baseline;
- simple rule extraction for a business process;
- a visual way to explain non-linear splits;
- a stepping stone before Random Forest or boosting.
Avoid relying on a single tree when:
- small data changes create very different trees;
- test score drops far below train score;
- the problem needs the accuracy and stability of ensembles.
Practical Debugging Checklist
| Symptom | Likely cause | Fix |
|---|---|---|
| Train score is high, test score is low | tree is too deep | lower max_depth, increase min_samples_leaf, try pruning |
| Tree has many tiny leaves | memorizing rare cases | increase min_samples_leaf |
| Feature importance feels misleading | correlated or high-cardinality features | verify with permutation importance |
| Rules are hard to read | tree is too large | train a smaller explanation tree, or summarize paths |
| Regression tree predictions look blocky | leaf averages create step outputs | compare with linear models, random forests, or gradient boosting |
Practice
- Change
min_samples_leaffrom3to1, then to10. What happens to leaves and test accuracy? - Change
criterionto"entropy". Does the tree choose the same first split? - Print
export_text()for a tree withmax_depth=2. Is it easier to explain? - Replace the Iris features with all four features. Does feature importance change?
- In the regression section, compare
DecisionTreeRegressorwith the linear regression lesson's baseline.
Pass Check
You are done when you can explain:
- a tree learns by choosing splits that make child nodes cleaner;
- deeper trees can memorize training data;
max_depth,min_samples_leaf, andccp_alphacontrol complexity;- feature importance is useful but not automatically causal;
- regression trees output leaf averages, so predictions are step-like.