11.4.2 Sequence Labeling Tasks
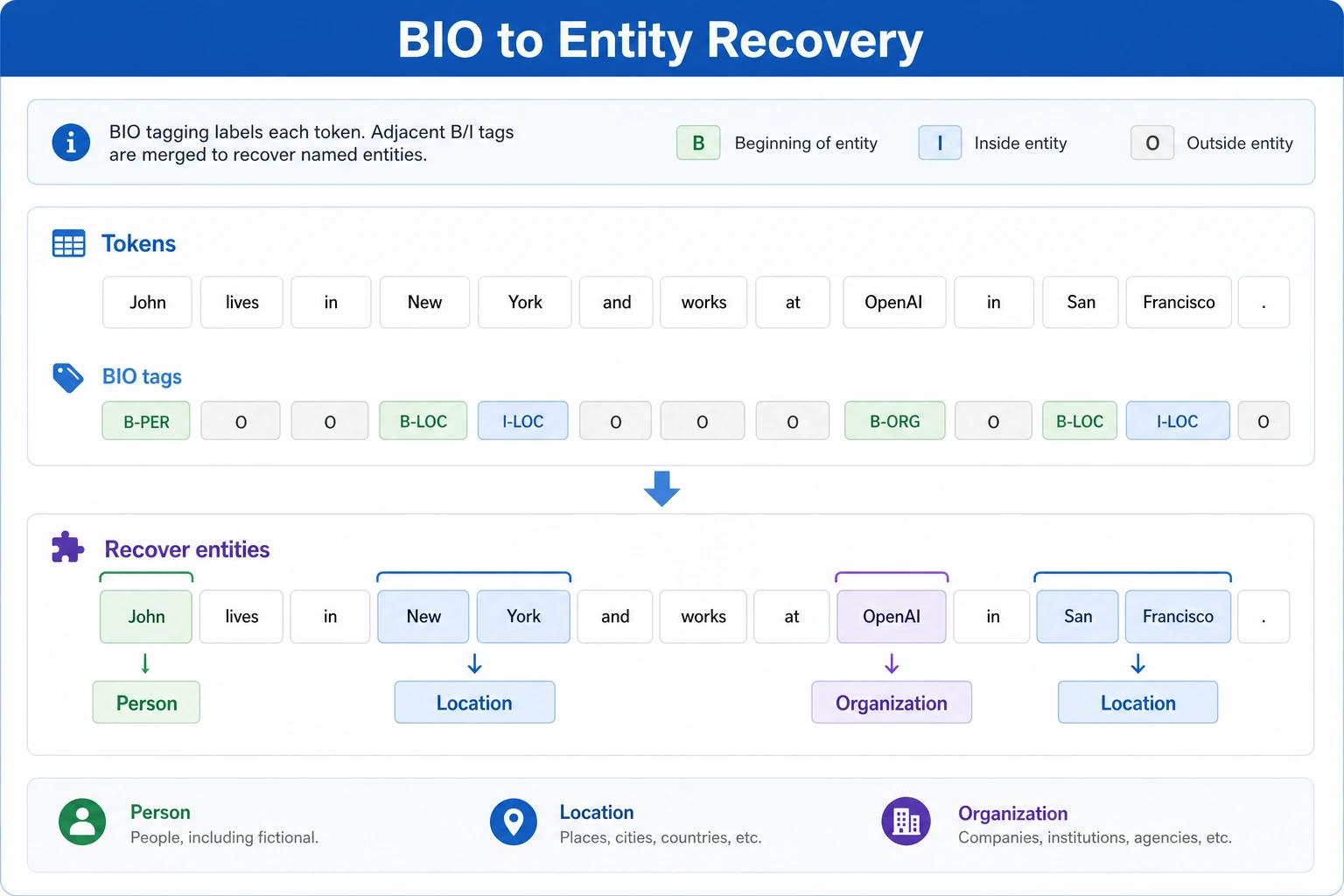
The output of text classification is usually:
- one label for the whole sentence
But the output of sequence labeling is more fine-grained:
- one label for each token
This step is very important because it moves NLP from “judging the whole sentence” to:
locating specific information inside the sentence.
From here, it becomes much more natural to move toward tasks like named entity recognition, information extraction, and slot filling.
Learning Objectives
- Understand the fundamental difference between sequence labeling and sentence-level classification
- Understand why label schemes such as BIO / BIOES are commonly used
- Use a runnable example to understand token-level labeling
- Build the connection between sequence labeling and information extraction tasks
What Problem Is Sequence Labeling Solving?
It is not just deciding “what kind of sentence this is,” but “which part of the sentence is what”
For example, the sentence:
- “Zhang San works at Peking University”
If you do text classification, you might only output:
- This is a sentence about a person and a location
But sequence labeling cares more about:
Zhang Sanis a person namePeking Universityis an organization name
Why is this important?
Because many real-world applications are not satisfied with sentence-level understanding. They care more about:
- person names
- addresses
- organization names
- amounts
- time expressions
That is, the positions and boundaries of these specific spans.
An analogy
Text classification is like putting a label on an entire article. Sequence labeling is like using a highlighter to circle important parts in the sentence.
Why Is the Output Usually Token-Based?
Because entities are continuous spans
Many pieces of information we want to extract are not single words, but a continuous span. For example:
Shanghai Jiao Tong UniversityJune 1, 2025
Token-level labels can express boundaries
That is why common label schemes do not simply write:
- PERSON
- LOCATION
Instead, they write:
B-PERI-PERO
The intuition behind BIO
B-: beginning of an entityI-: inside an entityO: not part of any entity
This lets the system distinguish more clearly:
- where an entity starts
- where it ends
First Run a Minimal BIO Labeling Example
tokens = ["Zhang San", "works at", "Peking", "University", "today"]
tags = ["B-PER", "O", "B-ORG", "I-ORG", "O"]
for tok, tag in zip(tokens, tags):
print(tok, tag)
Expected output:
Zhang San B-PER
works at O
Peking B-ORG
University I-ORG
today O
The token list and the tag list have the same length. That one-to-one alignment is the first thing to verify in every sequence-labeling dataset.
What is the most important thing in this example?
It shows you:
- a sequence input
- a corresponding sequence output
This is the most essential form of sequence labeling:
Input a sequence of tokens, output a sequence of labels of the same length.
Why are Peking University labeled as B-ORG / I-ORG?
Because the goal here is to express:
- this is one continuous entity
not two separate entities.
Recovering Entities from a Label Sequence
The following example recovers entity spans from token + BIO labels.
tokens = ["Zhang San", "works at", "Peking", "University", "today"]
tags = ["B-PER", "O", "B-ORG", "I-ORG", "O"]
def decode_entities(tokens, tags):
entities = []
current_tokens = []
current_type = None
for token, tag in zip(tokens, tags):
if tag == "O":
if current_tokens:
entities.append((" ".join(current_tokens), current_type))
current_tokens = []
current_type = None
continue
prefix, entity_type = tag.split("-", 1)
if prefix == "B":
if current_tokens:
entities.append((" ".join(current_tokens), current_type))
current_tokens = [token]
current_type = entity_type
elif prefix == "I" and current_type == entity_type:
current_tokens.append(token)
else:
# If the label is invalid, simply cut off and restart
if current_tokens:
entities.append((" ".join(current_tokens), current_type))
current_tokens = [token]
current_type = entity_type
if current_tokens:
entities.append((" ".join(current_tokens), current_type))
return entities
print(decode_entities(tokens, tags))
Expected output:
[('Zhang San', 'PER'), ('Peking University', 'ORG')]
This is the step that turns token-level labels into the project output people actually use: entity text plus entity type.
Why is this code important?
Because it connects the “labeling task” with the “extraction result.” In real systems, what we usually care about is not the labels themselves, but:
- entity spans
- entity types
What Is the Relationship Between Sequence Labeling and Information Extraction?
NER is a typical sequence labeling task
The most classic example is:
- named entity recognition
But it is not only used for NER
It can also be used for:
- slot filling
- keyword extraction
- event trigger identification
So it is a “foundational skill” for information extraction
Many extraction systems become more complex later, but the most basic first step is often still:
- mark the key spans first
Common Pitfalls
Mistake 1: Treating sequence labeling like ordinary classification
The biggest difference from sentence-level classification is:
- the output is sequence-aligned
Mistake 2: Only looking at labels and ignoring boundary recovery
Real systems care more about the final extracted entity spans, not the label table itself.
Mistake 3: Designing the label scheme casually
If the label design is messy, both the model and the evaluation will become messy too.
Summary
The most important takeaway from this lesson is to build one core intuition:
The core of sequence labeling is to assign labels to each token in the input sequence, so that the key spans and boundaries inside the sentence can be recovered.
Once this intuition is solid, it will be much smoother to learn NER, BiLSTM+CRF, and information extraction projects later.
Exercises
- Add another time entity to the example, such as
2025, and write a BIO label sequence yourself. - Why is the key role of the BIO label scheme to express entity boundaries?
- Explain in your own words: what is the biggest difference between sequence labeling and text classification?
- Think about this: if an invalid
I-XXXappears in the label sequence, how should the system handle it more robustly?