11.4.1 Sequence Labeling Roadmap: One Label per Token
Sequence labeling predicts one label for each token. NER, word segmentation, part-of-speech tagging, and slot filling all use this idea.
See the Label Path First
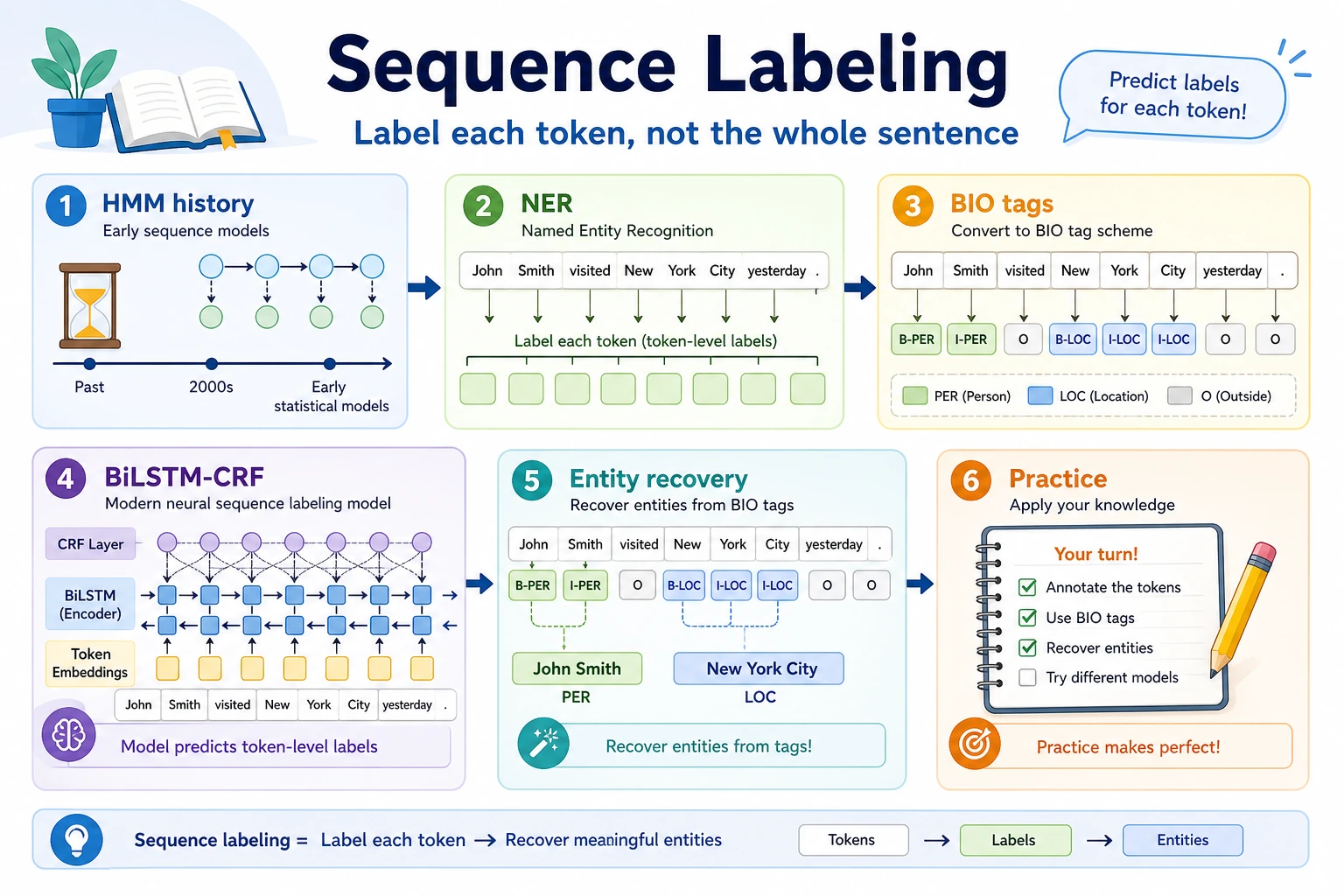
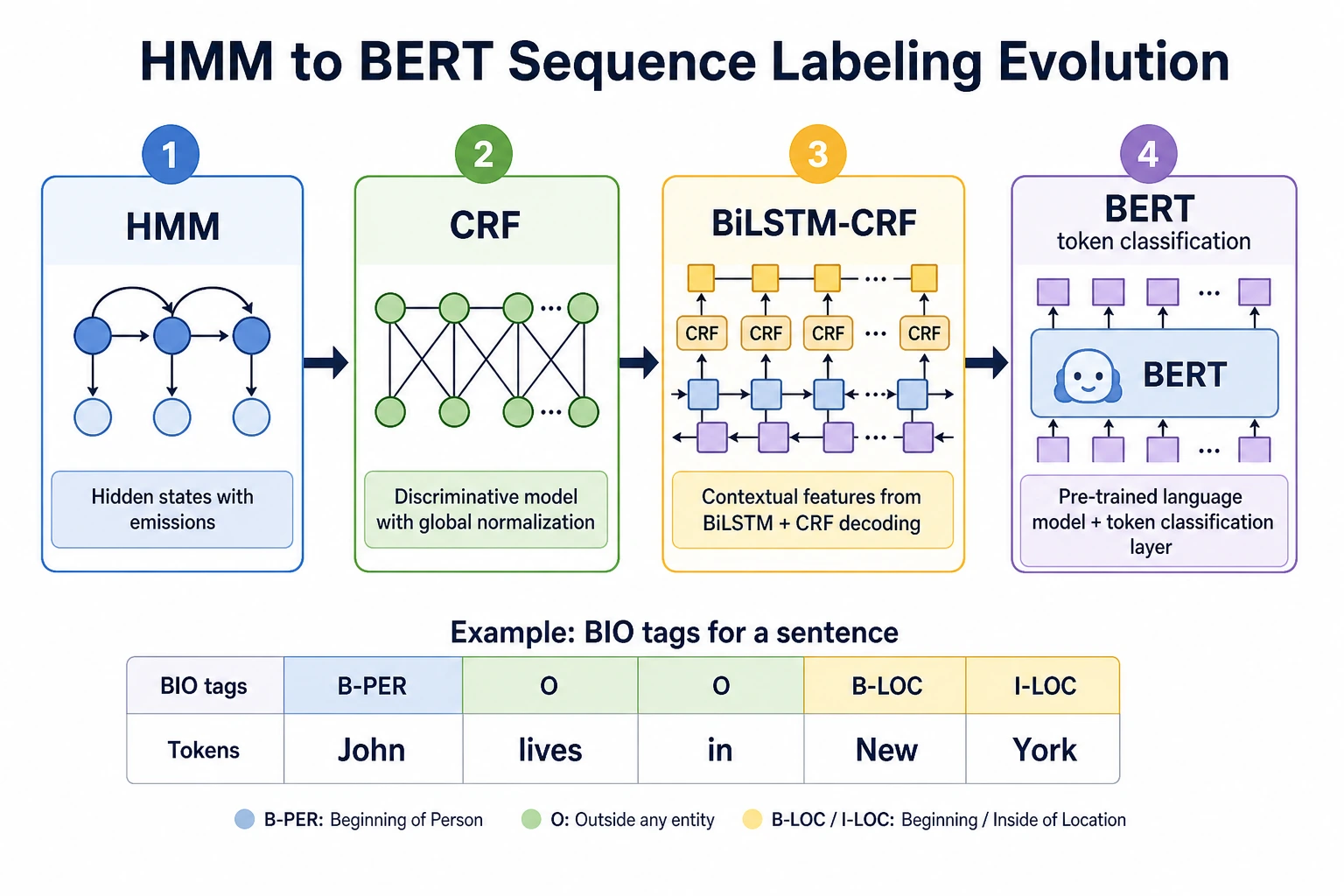
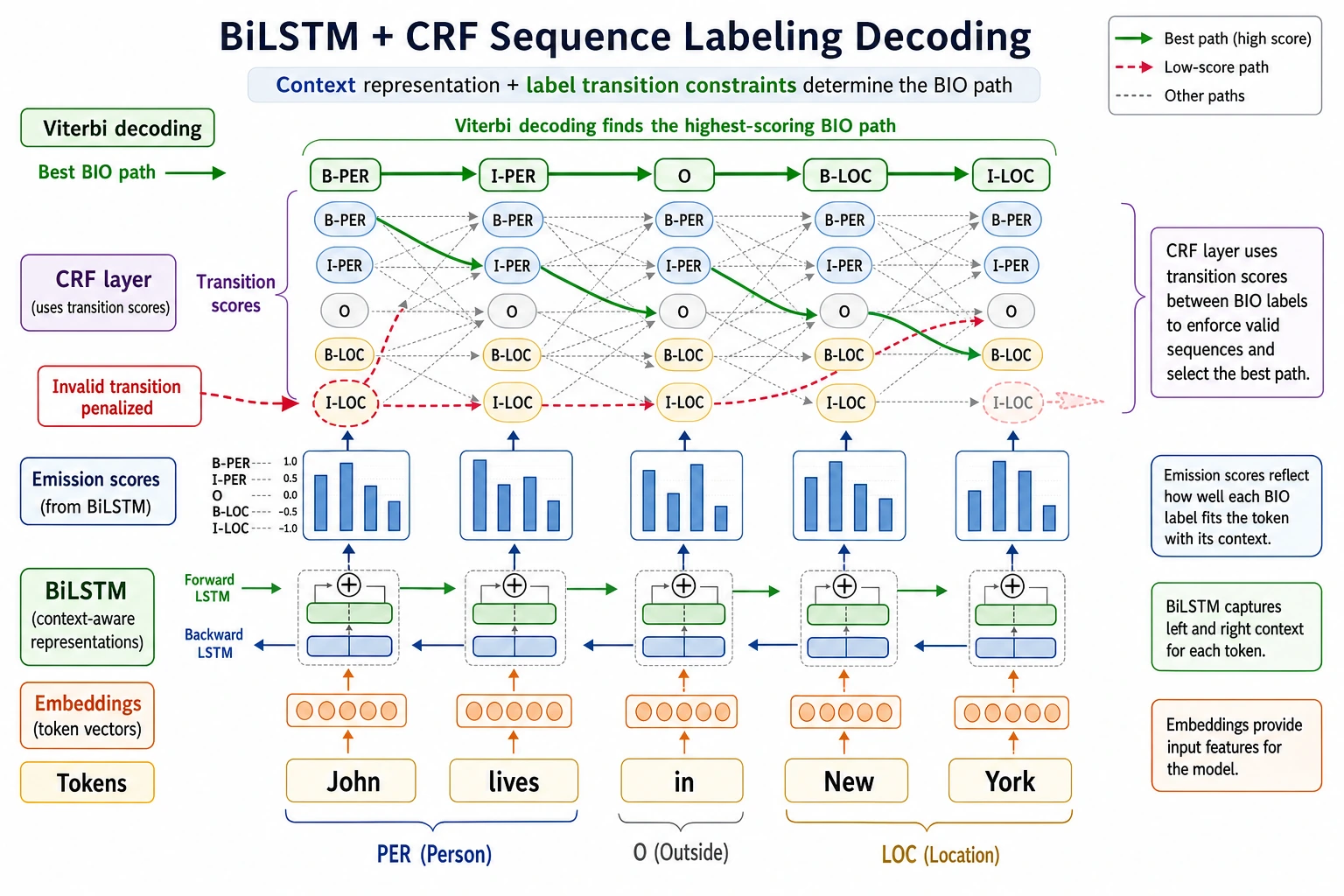
The key output is not one sentence label, but aligned token-level tags such as B-PER, I-PER, and O.
Run a BIO Tag Check
tokens = ["Ada", "Lovelace", "wrote", "notes"]
tags = ["B-PER", "I-PER", "O", "O"]
for token, tag in zip(tokens, tags):
print(token, tag)
Expected output:
Ada B-PER
Lovelace I-PER
wrote O
notes O
If tokenization changes, labels must stay aligned. Many sequence-labeling bugs are alignment bugs.
Learn in This Order
| Step | Read | Practice Output |
|---|---|---|
| 1 | NER and BIO | Create token-level labels and entity spans |
| 2 | HMM/CRF history | Understand sequence constraints and label transitions |
| 3 | BiLSTM-CRF | Connect contextual features with valid label paths |
| 4 | Project practice | Evaluate precision, recall, F1, boundary errors |
Pass Check
You pass this chapter when you can inspect token/tag alignment and explain one boundary error or invalid tag transition.