6.2.5 nn.Module
nn.Module is how PyTorch packages layers, parameters, forward logic, and training/evaluation mode into one model object. This section upgrades the hand-written parameters from autograd into reusable model classes.
Learning Objectives
- Use
nn.Linearand read its parameter shapes. - Build simple models with
nn.Sequential. - Write a custom
nn.Modulewith__init__()andforward(). - Inspect
named_parameters()andstate_dict(). - Understand what
model.train()andmodel.eval()actually switch.
Look at the Model Container
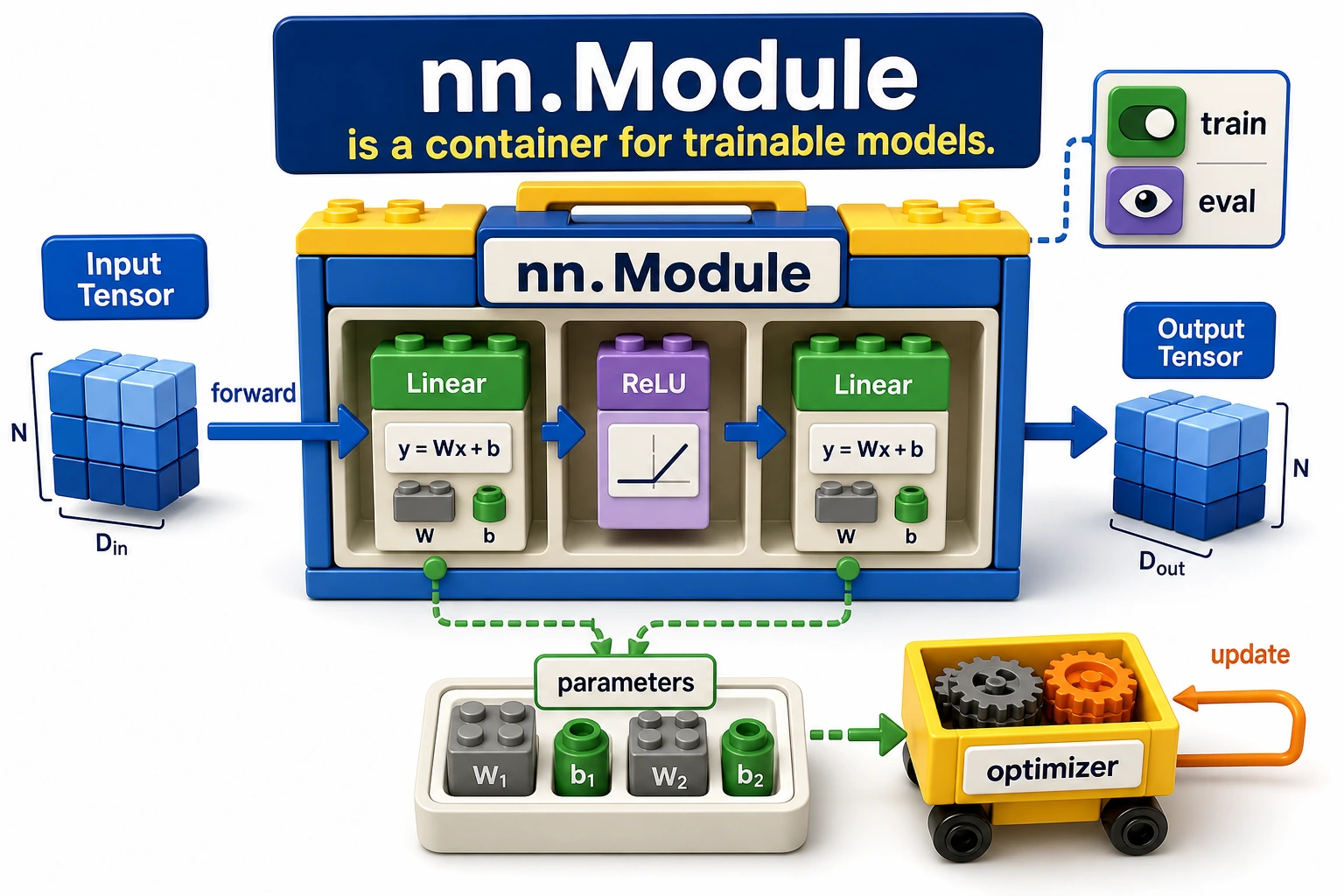
Think of nn.Module as a model container:
layers + parameters + forward logic + mode state -> one model object
The optimizer can then receive model.parameters() without needing to know how many layers you wrote.
From Manual Weights to nn.Linear
In the previous sections, you saw the operation:
logits = X @ W + b
nn.Linear(in_features, out_features) packages the same idea as a trainable layer.
import torch
from torch import nn
layer = nn.Linear(3, 2)
with torch.no_grad():
layer.weight.copy_(
torch.tensor(
[
[0.1, 0.2, 0.3],
[-0.1, 0.4, 0.2],
]
)
)
layer.bias.copy_(torch.tensor([0.01, -0.02]))
x = torch.tensor([[1.0, 2.0, 3.0]])
y = layer(x)
print("linear_lab")
print("input shape:", tuple(x.shape))
print("weight shape:", tuple(layer.weight.shape))
print("bias shape:", tuple(layer.bias.shape))
print("output:", torch.round(y * 100) / 100)
Expected output:
linear_lab
input shape: (1, 3)
weight shape: (2, 3)
bias shape: (2,)
output: tensor([[1.4100, 1.2800]], grad_fn=<DivBackward0>)
Important shape rule:
- input:
[batch, in_features] - weight:
[out_features, in_features] - output:
[batch, out_features]
The printed grad_fn means the output is connected to an autograd graph.
Build a Simple Network with nn.Sequential
Use nn.Sequential when data flows through layers in a straight line.
import torch
from torch import nn
torch.manual_seed(11)
model = nn.Sequential(
nn.Linear(3, 4),
nn.ReLU(),
nn.Linear(4, 2),
)
batch = torch.randn(5, 3)
logits = model(batch)
print("logits shape:", tuple(logits.shape))
Expected output:
logits shape: (5, 2)
Read the model:
[batch, 3] -> Linear(3, 4) -> ReLU -> Linear(4, 2) -> [batch, 2]
That is already a small multilayer perceptron.
Write a Custom nn.Module
Custom modules are the normal style for real projects because they can hold named submodules, branching logic, reusable helper methods, and clearer debugging hooks.
import torch
from torch import nn
class TinyClassifier(nn.Module):
def __init__(self, in_features=3, hidden=4, classes=2):
super().__init__()
self.net = nn.Sequential(
nn.Linear(in_features, hidden),
nn.ReLU(),
nn.Linear(hidden, classes),
)
def forward(self, x):
return self.net(x)
torch.manual_seed(11)
model = TinyClassifier()
batch = torch.randn(5, 3)
logits = model(batch)
print("module_lab")
print("logits shape:", tuple(logits.shape))
for name, param in model.named_parameters():
print(name, tuple(param.shape))
print("state keys:", list(model.state_dict().keys()))
Expected output:
module_lab
logits shape: (5, 2)
net.0.weight (4, 3)
net.0.bias (4,)
net.2.weight (2, 4)
net.2.bias (2,)
state keys: ['net.0.weight', 'net.0.bias', 'net.2.weight', 'net.2.bias']
Responsibilities:
| Method or API | Responsibility |
|---|---|
__init__() | create layers and submodules |
forward() | describe how input becomes output |
parameters() | return learnable parameters for the optimizer |
named_parameters() | expose parameter names and shapes for debugging |
state_dict() | expose tensors that can be saved and loaded |
Keep training logic out of forward(). Loss, backward(), and optimizer.step() belong to the training loop, not to the model definition.
train() and eval() Are Mode Switches
model.train() does not run the training loop, and model.eval() does not run validation. They switch the behavior of layers such as Dropout and BatchNorm.
Run this example:
import torch
from torch import nn
class DropoutProbe(nn.Module):
def __init__(self):
super().__init__()
self.dropout = nn.Dropout(p=0.5)
def forward(self, x):
return self.dropout(x)
probe = DropoutProbe()
sample = torch.ones(6)
torch.manual_seed(3)
probe.train()
train_a = probe(sample)
train_b = probe(sample)
probe.eval()
eval_a = probe(sample)
eval_b = probe(sample)
print("mode_lab")
print("train outputs equal:", torch.equal(train_a, train_b))
print("eval outputs equal:", torch.equal(eval_a, eval_b))
print("eval output:", eval_a)
Expected output:
mode_lab
train outputs equal: False
eval outputs equal: True
eval output: tensor([1., 1., 1., 1., 1., 1.])
Practical habit:
model.train() # before training batches
model.eval() # before validation or prediction
For validation, combine it with torch.no_grad():
model.eval()
with torch.no_grad():
logits = model(batch)
Mini Project: Train a Score Predictor
This example uses two features and one regression target:
- study hours per week;
- practice problems completed per week;
- predicted score.
The target is divided by 100 so the optimization is stable on this tiny dataset.
import torch
from torch import nn
class ScorePredictor(nn.Module):
def __init__(self):
super().__init__()
self.net = nn.Sequential(
nn.Linear(2, 16),
nn.ReLU(),
nn.Linear(16, 1),
)
def forward(self, x):
return self.net(x)
torch.manual_seed(42)
X = torch.tensor(
[
[2.0, 1.0],
[3.0, 2.0],
[4.0, 3.0],
[5.0, 5.0],
[6.0, 6.0],
[7.0, 8.0],
]
)
y = torch.tensor(
[
[55.0],
[60.0],
[68.0],
[78.0],
[85.0],
[92.0],
]
) / 100.0
model = ScorePredictor()
loss_fn = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=0.03)
print("training_lab")
for epoch in range(401):
pred = model(X)
loss = loss_fn(pred, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if epoch % 100 == 0:
print(f"epoch={epoch:3d} loss={loss.item():.4f}")
model.eval()
with torch.no_grad():
test = torch.tensor([[6.5, 7.0]])
pred_score = model(test).item() * 100
print("predicted score:", round(pred_score, 2))
Expected output:
training_lab
epoch= 0 loss=0.4672
epoch=100 loss=0.0003
epoch=200 loss=0.0001
epoch=300 loss=0.0001
epoch=400 loss=0.0001
predicted score: 89.31
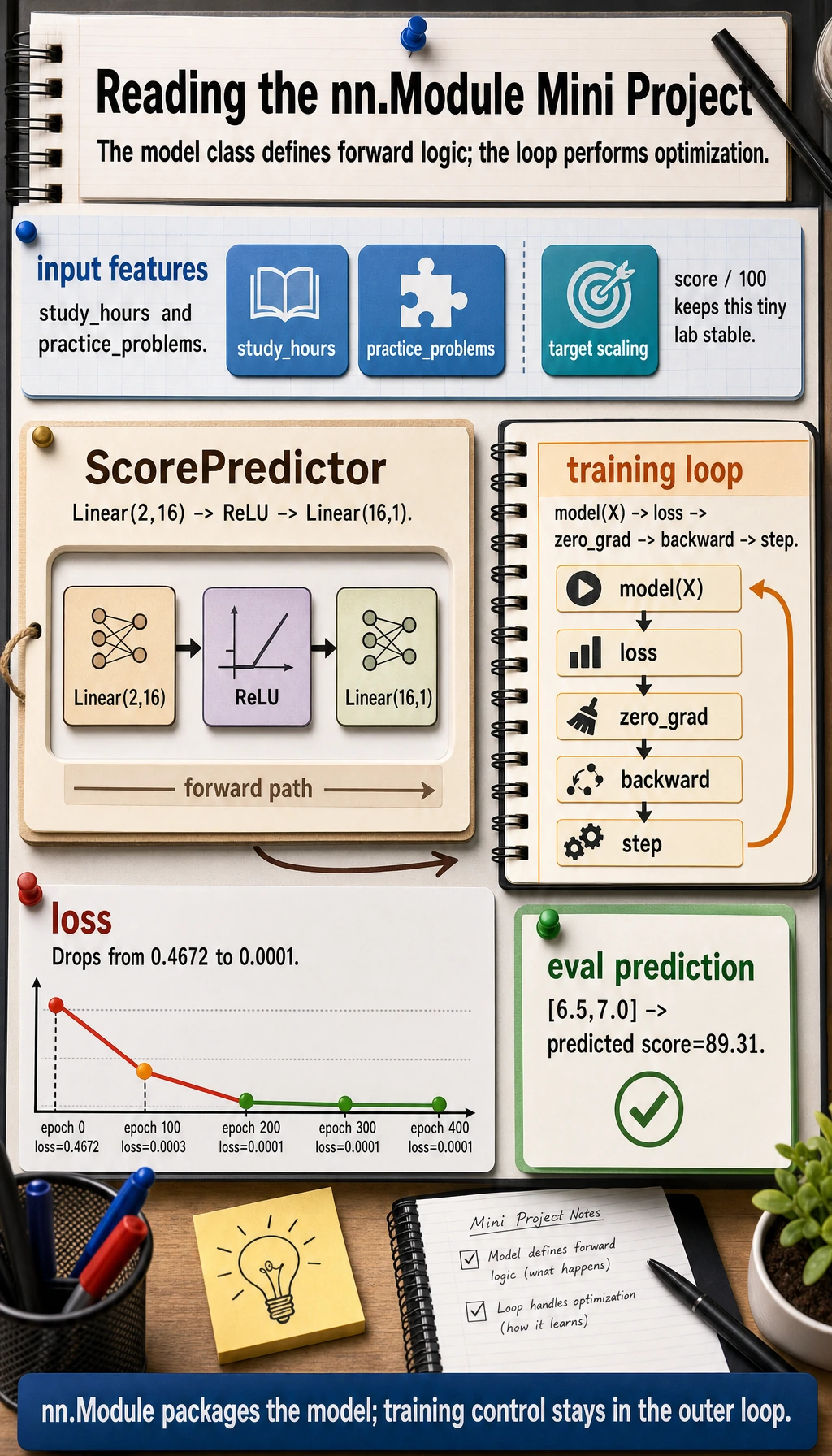

This is now a complete miniature PyTorch model:
data -> model -> loss -> zero_grad -> backward -> optimizer.step -> eval prediction
Sequential or Custom Module?
| Situation | Good choice |
|---|---|
| simple straight-line stack | nn.Sequential |
| multiple inputs or outputs | custom nn.Module |
| skip connections or branches | custom nn.Module |
| reusable components | custom nn.Module |
| you need clearer parameter names | custom nn.Module |
In real deep learning projects, custom modules are more common because architectures quickly become more than a straight line.
Common Mistakes
| Mistake | Why it hurts | Fix |
|---|---|---|
creating layers inside forward() | new parameters are created on every call and may not be optimized correctly | define layers in __init__() |
putting loss and optimizer logic inside forward() | mixes model definition with training control | keep forward() as input-to-output only |
forgetting super().__init__() | submodules may not register correctly | call it first in __init__() |
| not checking parameter names | hard to debug frozen or missing layers | print named_parameters() |
forgetting eval() for validation | Dropout/BatchNorm behave like training | call model.eval() before validation |
Exercises
- Change the hidden size in
ScorePredictorfrom16to4and32. How does the loss change? - Remove
ReLU(). Does the model still learn this tiny regression task? Why might deeper nonlinear tasks need it? - Print
model.state_dict()keys and shapes. Which tensors would be saved in a checkpoint? - Add
nn.Dropout(p=0.2)after ReLU, then compare predictions intrain()andeval()modes.
Key Takeaways
nn.Modulemanages layers, parameters, forward logic, and mode state together.forward()should describe data flow, not the training loop.model.parameters()is what connects the model to the optimizer.state_dict()is the standard checkpoint interface.train()andeval()switch layer behavior; they do not run loops by themselves.